Hi Dorian,
Welcome to NeuroStars!
That is a beauty of the git submodules which DataLad uses for its sub-datasets mechanism. If you place your DICOMs into a separate subdataset at sourcedata/, number of files in that subdataset should not have direct effect on its parent (BIDS) dataset. Moreover, you are typically not interested in sourcedata/ while working on already existing BIDS dataset, so it will allow to install BIDS dataset without all the dicoms etc. Then I think you will not get your millions of files in BIDS dataset.
For your sourcedata/ dataset – Git indeed becomes slower while working with growing number of files. Depending on your workflow you could have just .tar.gz’ed each series of DICOMs into its own .tar.gz thus minimizing number of files in the repo. E.g. we are doing that within ReproIn heuristic of HeuDiConv, which can work on tarballs of DICOMs.
10k was kinda a very low bound, there are datasets with >100k files which are still “functional”. With DICOMs dataset I do not expect you to do much/often, so I think it will be just fine. But also you could try (I haven’t yet) a relatively new feature of git – split index, see Git - git-update-index Documentation which should be of help in such cases.
not now, and not sure if ever (for SGE specifically). There is GitHub - datalad/datalad-htcondor: Remote code execution for DataLad via HTCondor · GitHub for local htcondor powered execution, and we are also developing reproman run which could then be used locally or you could schedule execution on remote service. See https://reproman.readthedocs.io/en/latest/execute.html for the basic docs about it. There is a PR with an example I reference below as well. There (in ReproMan) we currently provide support for PBS (torque) and Condor, no SGE. We hope to provide submitter for SLURM some time soon (SLURM submitter · Issue #484 · ReproNim/reproman · GitHub). Not sure if anyone in our team(s) would work on SGE though – we have used it only briefly in the past before quickly running away from it ![]() If you really need it, you could craft a submitter based on others, see e.g. one for Condor .
If you really need it, you could craft a submitter based on others, see e.g. one for Condor .
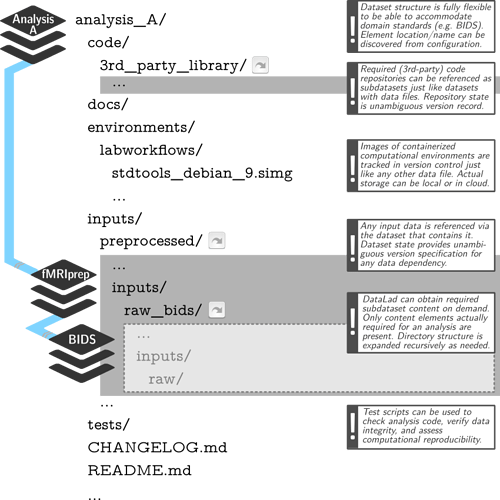
Well, if you were to follow YODA principles more closely, then BIDS dataset should not contain derivatives. You could organize your YODA study dataset to contain BIDS and derivatives subdatasets. Each derivative dataset should then “contain” BIDS dataset(s) it used. Those could be installed “temporarily and efficiently” (e.g., via datalad install --reckless --source ../bids raw_bids or relying on CoW functionality of filesystems like BTRFS). Each derivative dataset then could have raw_bids/ subdataset where BIDS dataset would be installed, and thus be self-sufficient and have all tracking information. See e.g.
If that is “too much”, then indeed at least making those to be subdatasets under
derivatives/ of BIDS dataset - would be the least of the YODA sins Sure, why not? ![]() it will initiate the dataset at the
it will initiate the dataset at the /data/studies level so you could add all the subdatasets to be contained within it.
I think it generally should be fine, but you might like to have some text files under git-annex control, e.g. if they contain possibly sensitive information, e.g. the _scans.tsv files could contain exact scanning date. That is why in heudiconv we explicitly list those to go under git-annex: heudiconv/heudiconv/external/dlad.py at master · nipy/heudiconv · GitHub .
If you install datalad-neuroimaging extension, then it would provide cfg_bids procedure, so you could use datalad create -c bids which establishes a bit less generous specification – only some top level files (README, CHANGES) go to git, the rest to git-annex.
Another point, with sensitive information in mind, you might like to start using --fake-dates option of datalad create for your DICOM and BIDS datasets if the dates when you add data to them are close to the original scanning date. That would make all git/git-annex commits start in the past and go at 1 sec interval between commits regardless when you datalad save ![]()
Oh hoh, I never found a peace of mind with ACL. git-annex needs additional work to provide lock down on ACL systems. Paranoid me also afraid of git-annex “shared” mode since it would then allows to introduce changes to tracked by it files . Having said all that, the easiest solution is just to have two clones (possibly from a “central” shared, possibly bare) with users having read-only access to the repository of the other user. I think it should work with ACL or just regular POSIX system permissions established at the level of the group having read permissions and user umasks not resetting group permissions (so being 022)
How would like them to be shared/exposed? And should it point to the license of the entrypoint project, or have licenses for all used/included components? Could be too difficult to assemble, besides for components installed via apt since all those packages must have licensing information in corresponding /usr/share/doc/<package>/copyright. Anyways – better file an issue with ReproNim/containers and we will see what could be done
That one specifically for submission of ISATAB files to data descriptor papers to Nature “Scientific Data”. You don’t need it to answer your question. You just need to extract (and possibly aggregate into super-datasets) metadata, after installing datalad-neuroimaging extension and enabling bids metadata extractor (git config -f .datalad/config --add datalad.metadata.nativetype bids). Then datalad aggregate-metadata should aggregate metadata. Then, though you can’t answer ‘which participant’, you could find an answer for “which T1w” I have across the collection of datasets I have:
(git-annex)lena:~/datalad[master]git
$> datalad -c datalad.search.index-egrep-documenttype=all search bids.type:T1w
search(ok): /home/yoh/datalad/dbic/QA/sourcedata/sub-amit/ses-20180508/anat/sub-amit_ses-20180508_acq-MPRAGE_T1w.dicom.tgz (file)
search(ok): /home/yoh/datalad/dbic/QA/sourcedata/sub-emmet/ses-20180508/anat/sub-emmet_ses-20180508_acq-MPRAGE_T1w.dicom.tgz (file)
search(ok): /home/yoh/datalad/dbic/QA/sourcedata/sub-emmet/ses-20180521/anat/sub-emmet_ses-20180521_acq-MPRAGE_T1w.dicom.tgz (file)
search(ok): /home/yoh/datalad/dbic/QA/sourcedata/sub-emmet/ses-20180531/anat/sub-emmet_ses-20180531_acq-MPRAGE_T1w.dicom.tgz (file)
search(ok): /home/yoh/datalad/dbic/QA/sourcedata/sub-qa/ses-20171030/anat/sub-qa_ses-20171030_acq-MPRAGE_T1w.dicom.tgz (file)
search(ok): /home/yoh/datalad/dbic/QA/sourcedata/sub-qa/ses-20171106/anat/sub-qa_ses-20171106_acq-MPRAGE_T1w.dicom.tgz (file)
search(ok): /home/yoh/datalad/dbic/QA/sourcedata/sub-qa/ses-20171113/anat/sub-qa_ses-20171113_acq-MPRAGE_T1w.dicom.tgz (file)
search(ok): /home/yoh/datalad/dbic/QA/sourcedata/sub-qa/ses-20171120/anat/sub-qa_ses-20171120_acq-MPRAGE_T1w.dicom.tgz (file)
search(ok): /home/yoh/datalad/dbic/QA/sourcedata/sub-qa/ses-20171127/anat/sub-qa_ses-20171127_acq-MPRAGE_T1w.dicom.tgz (file)
search(ok): /home/yoh/datalad/dbic/QA/sourcedata/sub-qa/ses-20171204/anat/sub-qa_ses-20171204_acq-MPRAGE_T1w.dicom.tgz (file)
With ongoing development of GitHub - datalad/datalad-metalad: Next generation metadata handling · GitHub which is intended to replace metadata handling in datalad, we hope to provide ways to answer “which subject” or “which dataset”.
oh, there is no single location. Exemplar use cases should eventually appear in documentation(s) and handbook. E.g.
- An automatically and computationally reproducible neuroimaging analysis from scratch — The DataLad Handbook
- ReproNim training for OHBM 2018 (no HPC, just
datalad run): Computational basis and ReproIn/DataLad: ReproIn/DataLad: A complete portable and reproducible fMRI study from scratch - a prototypical workflow which would use
datalad runorreproman runfor a typical BIDS dataset analysis: (yet to be finished) Initial sketch for the mriqc/fmriprep singularity based workflow by yarikoptic · Pull Request #438 · ReproNim/reproman · GitHub - just search GitHub for DATALAD RUNCMD to see how us/others used
datalad run
Overall – everything is still a moving target ![]()