-
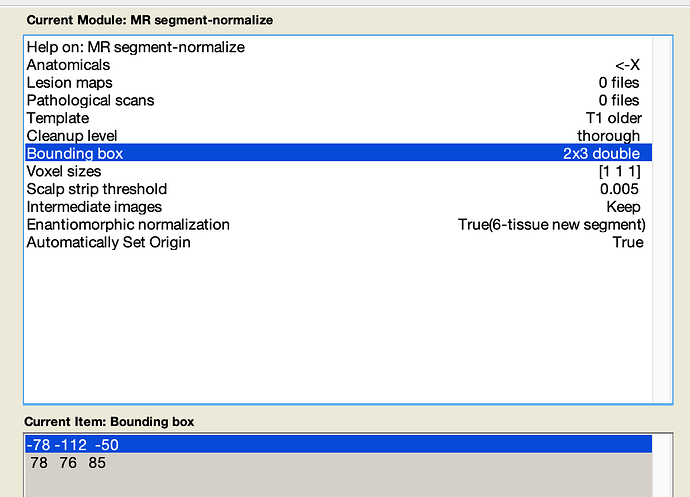
You can adjust the “bounding box” in the Clinical toolbox (and other SPM normalization scripts) to adjust the field-of-view of the resampled data, and you can adjust the voxel size to adjust the resolution. The initial values are the SPM defaults. As I recall, SPM images are typically replaced to an odd number of voxels in the left-right dimension (e.g. 181 vs 182), so the axis of symmetry runs through the center of voxels. You may want to specify why the 182x218x182 is specific for your application, rather than simply making images from all individuals consistent.
-
Yes, SPM normalizes brains to have be the size of an average brain, while the MNI template is larger than the average brain (as described in the original paper on the MNI template). Therefore, you really do not want to mix-and-match images warped to an MNI template with ANTS, FSL, etc. and from SPM. Further, note that the Clinical toolbox allows you to select between the “T1 older” and the “T1 younger” templates. The longer template matches the SPM defaults. The older template is based on neurologically healthy individuals age-matched to stroke patients. The older template demonstrates cerebral atrophy as one would expect. The intention is to maximize the match and minimize distortions. However, you should not mix and match participants aligned to different templates. The older template does show considerably larger ventricles, wider sulci, and the boundary of the cortex is clearly a bit different near the frontal cortex.
When working with clinical scans, please carefully visually inspect each solution. Most routines, including the clinical toolbox have issues when confronted with wide diploid space, which is associated with stroke risk factors. At some stage, I will release a new script for enantiomorphic normalization of stroke brains that leverages CAT12. In my experience, this provides the most robust solution to this problem, outperforming any solutions I have seen, including those I developed myself specifically for this problem.