Hi,
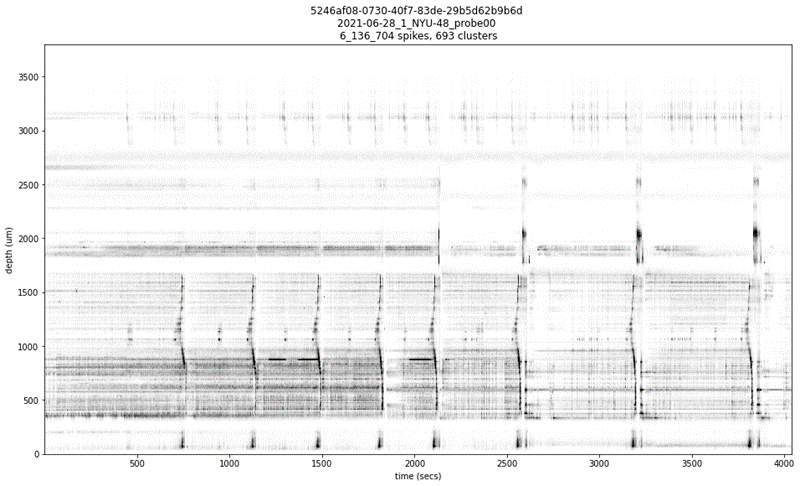
When exploring the dataset of ‘Reproducible Ephys Paper’, we found a ‘weird’ session with a lot of artifacts. The pid is ‘5246af08-0730-40f7-83de-29b5d62b9b6d’.
The minimum code example:
from one.api import ONE
from brainbox.io.one import SpikeSortingLoader
from ibllib.atlas import AllenAtlas
one = ONE(base_url='https://openalyx.internationalbrainlab.org')
ba = AllenAtlas()
pid = '5246af08-0730-40f7-83de-29b5d62b9b6d'
sl = SpikeSortingLoader(pid=pid, one=one, atlas=ba)
spikes, clusters, channels = sl.load_spike_sorting()
clusters = sl.merge_clusters(spikes, clusters, channels)
sl.raster(spikes)
The corresponding output is shown in Figure 1.
Figure 1
Even after selecting those ‘good’ neurons, we can still see similar patterns.
good_clusterIDs = clusters['cluster_id'][clusters['label'] == 1]
spikes_idx = np.isin(spikes['clusters'], good_clusterIDs)
spikes['times'] = spikes['times'][spikes_idx]
spikes['depths'] = spikes['depths'][spikes_idx]
spikes['clusters'] = spikes['clusters'][spikes_idx]
sl.raster(spikes)
The recording is also labelled as a ‘good’ insertion according to the ‘biorxiv_2022_05.csv’ file. We are a little bit worried that it will harm the conclusions researchers could draw from these data. Could you please have a check of it? Or would you think about dropping it from the public dataset?
Best wishes,
Fenying Zang
PhD student
Leiden University, The Netherlands
Hello,
Thank you for pointing this out.
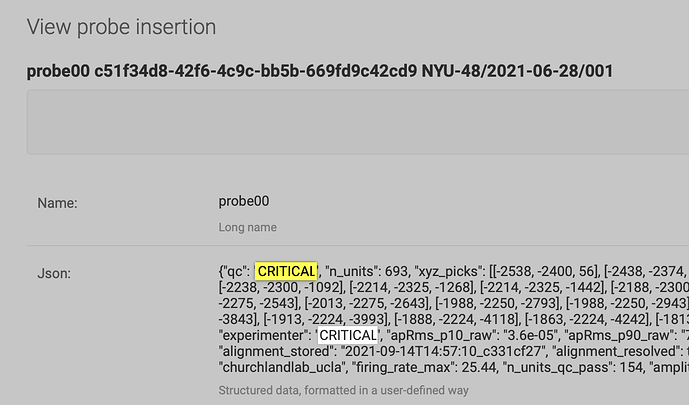
We indeed have flagged this recording as “CRITICAL”, cf qc field on Log in | Alyx
This dataset does not enter in the analysis. We will change the table to make it more clear, we have not updated it yet, thank you for your keen eye.
Kind regards
Many thanks for your quick reply. 
Best,
Fenying
You are welcome!
One note:
There was technically no issue with the experiment here, i.e. the hardware behaved fine, there is no noise nor no artefact (coming for example from ground loop).
Rather, the large disruption you see on the raster are “real” and due to epileptic seizures.
We obviously did not aim for any epileptiform activity ; but when doing experiment at a large scale (500+ recordings) this is likely to occur by chance.
Thanks for the information. It helps a lot because I have not done such kind of experiments yet. I will keep that in mind.
Have a nice day!
Best,
Fenying
Hi @Gaelle_Chapuis, chiming in here - good to know that’s the reason. Indeed this cannot be avoided, but it may make sense to exclude this from 1. the BWM and repro-ephys data releases and 2. from the list of ‘good’ sessions that are in the codebase now paper-reproducible-ephys/data_releases at master · int-brain-lab/paper-reproducible-ephys · GitHub? That way, we avoid studying epilepsy without intending to so do 
I hope I’m misunderstanding and 1. is already the case, which may mean that only 2. has an outdated spreadsheet. If that’s true, we’ll wait for the updated one (or see if we can recompute session-level QC on the public data ourselves).