Dear all,
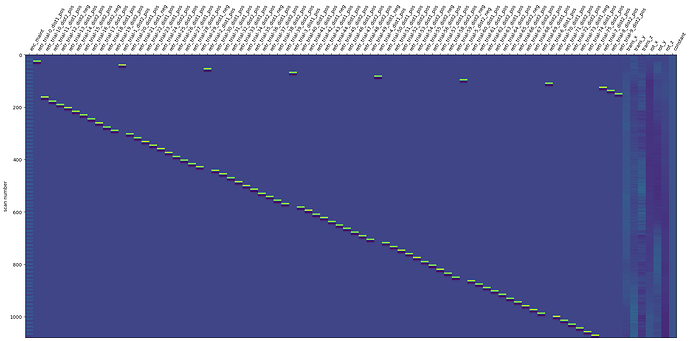
I am currently performing an fMRI task where participants do 72 trials of either task A or task B. I want to set up a decoder model to classify between A vs B. Following online info, I am first fitting a trial wise design matrix that looks like this (please ignore first column)
Then, I estimate the contrast against the implicit baseline for every trial
# %%
z_maps = []
trial_id = []
cond_id = []
pos_id = []
beta_names = first_level_model.design_matrices_[0].columns
for beta_indx, beta_name in enumerate(beta_names):
if "retr_trial" in beta_name:
contrast = np.zeros(len(beta_names))
contrast[beta_indx] = 1
z_maps.append(first_level_fit.compute_contrast(contrast))
trial_id.append(beta_name.split('_')[1])
cond_id.append(beta_name.split('_')[2])
pos_id.append(beta_name.split('_')[3])
Therefore, I have a list of z maps corresponding to contrasts, a list of the conditions and of the trials.
My mask consists of 515 voxels (in standard MNI 91 109 91 space).
To run the decoder, I do
decoder = Decoder(
estimator="svc",
mask = roi_mask,
cv=LeaveOneGroupOut(),
screening_percentile=20,
scoring="accuracy",
)
decoder.fit(pos_z_maps, pos_cond_id, groups=pos_trial_id)
However, I get the following warnings:
<ipython-input-129-c0f8ca558d72>:11: UserWarning: [NiftiMasker.fit] Generation of a mask has been requested (imgs != None) while a mask was given at masker creation. Given mask will be used.
decoder.fit(z_maps, cond_id, groups=trial_id)
<ipython-input-129-c0f8ca558d72>:11: FutureWarning: The 'zscore' strategy incorrectly uses population std to calculate sample zscores. The new strategy 'zscore_sample' corrects this behavior by using the sample std. In release 0.14.0, the 'zscore' option will be removed and using standardize=True will fall back to 'zscore_sample'.To avoid this warning, please use 'zscore_sample' instead.
decoder.fit(z_maps, cond_id, groups=trial_id)
<ipython-input-129-c0f8ca558d72>:11: MaskWarning: Brain mask is smaller than .5% of the size of the standard human brain. This object is probably not tuned to be used on such data.
decoder.fit(z_maps, cond_id, groups=trial_id)
<ipython-input-129-c0f8ca558d72>:11: UserWarning: screening_percentile set to '100' despite requesting 'screening_percentile=20'.
All elements in the mask will be included.
This usually occurs when the mask image is too small compared to full brain mask.
decoder.fit(z_maps, cond_id, groups=trial_id)
Why can I not use the mask as specified?
Thank you for your help