Hi,
Just wondering about the brain_mask created when running the dhcp-structural-pipeline.
It is created early on with BET on the T2 using:
$ bet N4/"T2"_rescaled.nii.gz segmentations/"T2"_brain.nii.gz -R -f 0.1 -m.
However, this brain_mask is often very large, containing lots of extra-cranial tissue (e.g. the neck). When I previously have worked with the DrawEM neonatal segmentation pipeline, I have copied my own brain_mask in to the location where I know the script will put it. Then I run the script, which then omits this step and uses the pre-existing brain mask.
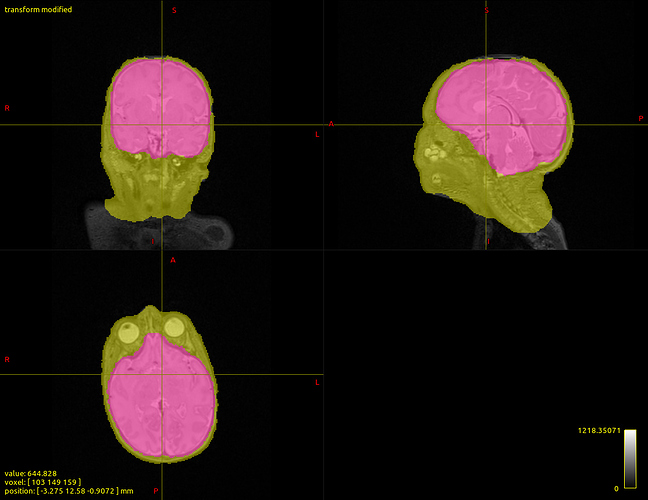
One example of my current data is provided, with the T2 with overlays of the dhcp-structural-pipeline brain extraction (yellow) as in the BET command above and my own brain extraction (magenta).
So, 2 questions
-
Would I benefit from a better brain_mask?
-
Where could I put it when running the dhcp-structural-pipeline via docker?
Cheers
Finn