Dear experts,
I recently tried to compare the tracts properties (FA, MD) of two groups. The tracts have been generated by mrtrix3. I first used AFQ to process, but it is difficult to covert .nii or .mif file to dt6 format which is acceptable for AFQ in Matlab. Then I found pyAFQ which is updated in recent years. However, there isn’t many examples about it. I wonder does it compute fiber properties between groups and make comparison?
Best wishes,
Thanks.
Hello,
I am not sure the statistics are available in PyAFQ, but you can definitely make the profiles in PyAFQ (Plotting tract profiles — AFQ 0.11 documentation) and should be able to do the group comparisons in DIPY (DIPY : Docs 1.5.0 - BUndle ANalytics (BUAN) framework).
Best,
Steven
Thanks, this is exactly what I am looking for. ![]()
Hi,
I meet another question. The download page of DIPY only provides the DIPY package in python. But how to install the DIPY BUAN framework, which seems a command in linux.
Thanks.
The command line tools should be installed in whatever environment you installed the python package.
Hi @Roger1219
When you install DIPY, all DIPY commandlines should automatically be installed including commandlines for BUAN framework. Of course, need to make sure you installed the package in your working environment as mentioned by @Steven
To test if commandlines are accessible through your terminal, you can try writing
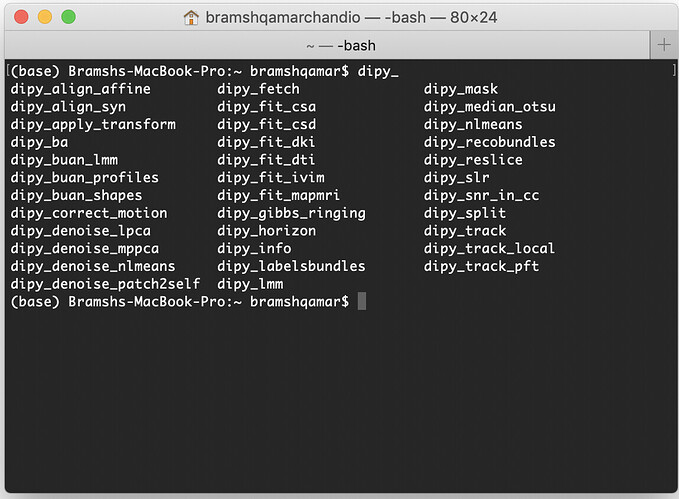
dipy_
and then press the ‘tab’ button from your keyboard. You should get a list of all available DIPY commandlines, something like this:
Or you can also try doing this from your terminal:
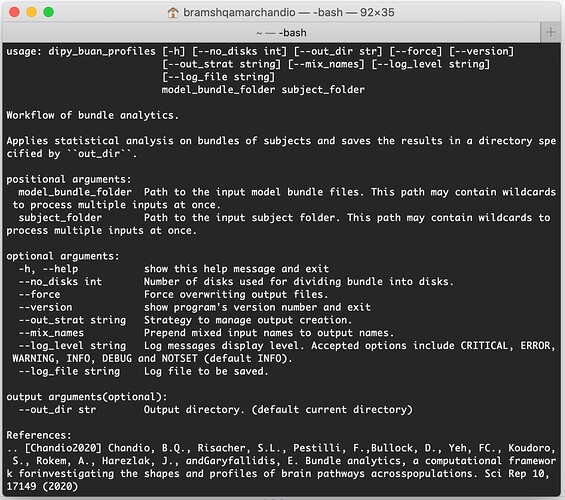
dipy_buan_profiles -h
It should show you a list of arguments required for the workflow/commandline and its documentation like this:
For the rest, you can follow this tutorial: https://dipy.org/documentation/1.5.0/interfaces/buan_flow/
You should be able to run BUAN on tracts extracted using mrtrix3. You may just need to format the directory/folder structure to match the one expected by BUAN workflows/commannlines as mentioned in the tutorial.
You can also ask DIPY specific questions here: https://gitter.im/dipy/dipy
Best,
Bramsh
Hello,
Thanks for your detailed response. I remembered that I installed DIPY on Conda environment and after I activate the env, I found the DIPY_ command.
Thanks to both of you and Steven!
Best,
Roger