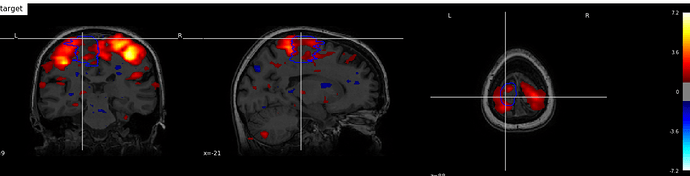
I have NIFTI files of anat, func and lesion areas for 1 subject. I made nilearn plot_stat_map for all nifty and have a result in image:
What i m trying to have is: crop regions of func (blue,red green) with lesion area, so the region of lesion should be clean of func inside countour, but with anat background. What is the way to do it?
Best Vyacheslav
What was my solution:
resampled_lesion_motor = nilearn.image.reample_to_image(motor, lesion) #resampling to same dim lesion and motor maps
motor_data = motor.get_data().copy()
motor_data[resampled_lesion_motor.get_data().astype(bool)] = 0 #zeroing region with get
motor_wo_lesion = image.new_img_like(motor, motor_data)
plotting.plot_epi(motor_wo_lesion)
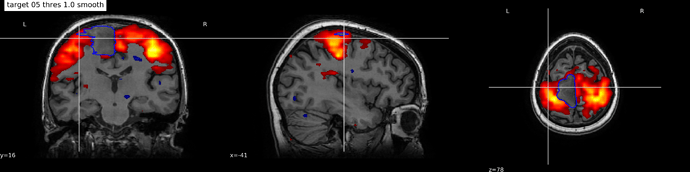
As it seen it on image motor( after ICA map) was cropped with lesion map.
sub-Bystrova_Motor_target-05_thres-1.0_smooth.png
1 Like