Summary of what happened:
Hi everyone, I am newish to fMRI analyses and trying to do a seed to voxel analysis in FSL FEAT (version 6.0.7.16). I preprocessed my data in fMRIprep and denoised it in Nilearn. The issue I am running into is that when I detrend the data in Nilearn, no clusters are detected in the FEAT analysis. I have confirmed the FEAT analysis does work and detects clusters when detrend=False in Nilearn. I have been playing around with adding the mask (in MNI space from fMRIprep Freesurfer outputs) in the design.fsf file, smoothing, and changing the thresholds, not of which seems to help. I have not applied a high_pass filter or sample_mask yet as I’m trying to troubleshoot this issue first. I would welcome any thoughts or suggestions on how to fix this either in Nilearn or FEAT parameters, below is the design.fsf file I have been using and the denoising code if helpful. Thank you in advance!
Command used (and if a helper script was used, a link to the helper script or the command generated):
PASTE CODE HERE
Version:
Environment (Docker, Singularity / Apptainer, custom installation):
Data formatted according to a validatable standard? Please provide the output of the validator:
PASTE VALIDATOR OUTPUT HERE
Relevant log outputs (up to 20 lines):
# FEAT version number
set fmri(version) 6.00
# Are we in MELODIC?
set fmri(inmelodic) 0
# Analysis level
# 1 : First-level analysis
# 2 : Higher-level analysis
set fmri(level) 1
# Which stages to run
# 0 : No first-level analysis (registration and/or group stats only)
# 7 : Full first-level analysis
# 1 : Pre-processing
# 2 : Statistics
set fmri(analysis) 2
# Use relative filenames
set fmri(relative_yn) 0
# Balloon help
set fmri(help_yn) 1
# Run Featwatcher
set fmri(featwatcher_yn) 1
# Cleanup first-level standard-space images
set fmri(sscleanup_yn) 0
# Output directory
set fmri(outputdir) "/Desktop/FEAT_test/FEAT_TEST2.feat"
# TR(s)
set fmri(tr) 2.500000
# Total volumes
set fmri(npts) 330
# Delete volumes
set fmri(ndelete) 0
# Perfusion tag/control order
set fmri(tagfirst) 1
# Number of first-level analyses
set fmri(multiple) 1
# Higher-level input type
# 1 : Inputs are lower-level FEAT directories
# 2 : Inputs are cope images from FEAT directories
set fmri(inputtype) 2
# Carry out pre-stats processing?
set fmri(filtering_yn) 0
# Brain/background threshold, %
set fmri(brain_thresh) 10
# Critical z for design efficiency calculation
set fmri(critical_z) 5.3
# Noise level
set fmri(noise) 0.66
# Noise AR(1)
set fmri(noisear) 0.34
# Motion correction
# 0 : None
# 1 : MCFLIRT
set fmri(mc) 0
# Spin-history (currently obsolete)
set fmri(sh_yn) 0
# B0 fieldmap unwarping?
set fmri(regunwarp_yn) 0
# GDC Test
set fmri(gdc) ""
# EPI dwell time (ms)
set fmri(dwell) 0.0
# EPI TE (ms)
set fmri(te) 0.0
# % Signal loss threshold
set fmri(signallossthresh) 10
# Unwarp direction
set fmri(unwarp_dir) y-
# Slice timing correction
# 0 : None
# 1 : Regular up (0, 1, 2, 3, ...)
# 2 : Regular down
# 3 : Use slice order file
# 4 : Use slice timings file
# 5 : Interleaved (0, 2, 4 ... 1, 3, 5 ... )
set fmri(st) 0
# Slice timings file
set fmri(st_file) ""
# BET brain extraction
set fmri(bet_yn) 0
# Spatial smoothing FWHM (mm)
set fmri(smooth) 0.0
# Intensity normalization
set fmri(norm_yn) 0
# Perfusion subtraction
set fmri(perfsub_yn) 0
# Highpass temporal filtering
set fmri(temphp_yn) 0
# Lowpass temporal filtering
set fmri(templp_yn) 0
# MELODIC ICA data exploration
set fmri(melodic_yn) 0
# Carry out main stats?
set fmri(stats_yn) 1
# Carry out prewhitening?
set fmri(prewhiten_yn) 0
# Add motion parameters to model
# 0 : No
# 1 : Yes
set fmri(motionevs) 0
set fmri(motionevsbeta) ""
set fmri(scriptevsbeta) ""
# Robust outlier detection in FLAME?
set fmri(robust_yn) 0
# Higher-level modelling
# 3 : Fixed effects
# 0 : Mixed Effects: Simple OLS
# 2 : Mixed Effects: FLAME 1
# 1 : Mixed Effects: FLAME 1+2
set fmri(mixed_yn) 2
# Higher-level permutations
set fmri(randomisePermutations) 5000
# Number of EVs
set fmri(evs_orig) 1
set fmri(evs_real) 1
set fmri(evs_vox) 0
# Number of contrasts
set fmri(ncon_orig) 1
set fmri(ncon_real) 1
# Number of F-tests
set fmri(nftests_orig) 0
set fmri(nftests_real) 0
# Add constant column to design matrix? (obsolete)
set fmri(constcol) 0
# Carry out post-stats steps?
set fmri(poststats_yn) 1
# Pre-threshold masking?
set fmri(threshmask) ""
# Thresholding
# 0 : None
# 1 : Uncorrected
# 2 : Voxel
# 3 : Cluster
set fmri(thresh) 3
# P threshold
set fmri(prob_thresh) 0.05
# Z threshold
set fmri(z_thresh) 1.5
# Z min/max for colour rendering
# 0 : Use actual Z min/max
# 1 : Use preset Z min/max
set fmri(zdisplay) 0
# Z min in colour rendering
set fmri(zmin) 2
# Z max in colour rendering
set fmri(zmax) 8
# Colour rendering type
# 0 : Solid blobs
# 1 : Transparent blobs
set fmri(rendertype) 1
# Background image for higher-level stats overlays
# 1 : Mean highres
# 2 : First highres
# 3 : Mean functional
# 4 : First functional
# 5 : Standard space template
set fmri(bgimage) 1
# Create time series plots
set fmri(tsplot_yn) 1
# Registration to initial structural
set fmri(reginitial_highres_yn) 0
# Search space for registration to initial structural
# 0 : No search
# 90 : Normal search
# 180 : Full search
set fmri(reginitial_highres_search) 90
# Degrees of Freedom for registration to initial structural
set fmri(reginitial_highres_dof) 3
# Registration to main structural
set fmri(reghighres_yn) 0
# Search space for registration to main structural
# 0 : No search
# 90 : Normal search
# 180 : Full search
set fmri(reghighres_search) 90
# Degrees of Freedom for registration to main structural
set fmri(reghighres_dof) BBR
# Registration to standard image?
set fmri(regstandard_yn) 0
# Use alternate reference images?
set fmri(alternateReference_yn) 0
# Standard image
set fmri(regstandard) "/Applications/fsl/data/standard/MNI152_T1_2mm_brain"
# Search space for registration to standard space
# 0 : No search
# 90 : Normal search
# 180 : Full search
set fmri(regstandard_search) 90
# Degrees of Freedom for registration to standard space
set fmri(regstandard_dof) 12
# Do nonlinear registration from structural to standard space?
set fmri(regstandard_nonlinear_yn) 0
# Control nonlinear warp field resolution
set fmri(regstandard_nonlinear_warpres) 10
# High pass filter cutoff
set fmri(paradigm_hp) 0
# Total voxels
set fmri(totalVoxels) 363127050
set fmri(fnirt_config) "T1_2_MNI152_2mm"
# Number of lower-level copes feeding into higher-level analysis
set fmri(ncopeinputs) 0
# 4D AVW data or FEAT directory (1)
set feat_files(1) "/Desktop/FEAT_test/sub-MC002/ses-01/func/sub-MC002_ses-01_denoise_24HMP_WMCSF_GSR_detrend2.nii.gz"
# Add confound EVs text file
set fmri(confoundevs) 0
# Session's alternate reference image for analysis 1
set alt_ex_func(1) "/Desktop/FEAT_test/sub-MC002/anat/sub-MC002_space-MNIPediatricAsym_cohort-6_res-2_desc-preproc_T1w.nii.gz"
# EV 1 title
set fmri(evtitle1) "Occ"
# Basic waveform shape (EV 1)
# 0 : Square
# 1 : Sinusoid
# 2 : Custom (1 entry per volume)
# 3 : Custom (3 column format)
# 4 : Interaction
# 10 : Empty (all zeros)
set fmri(shape1) 2
# Convolution (EV 1)
# 0 : None
# 1 : Gaussian
# 2 : Gamma
# 3 : Double-Gamma HRF
# 4 : Gamma basis functions
# 5 : Sine basis functions
# 6 : FIR basis functions
# 8 : Alternate Double-Gamma
set fmri(convolve1) 0
# Convolve phase (EV 1)
set fmri(convolve_phase1) 0
# Apply temporal filtering (EV 1)
set fmri(tempfilt_yn1) 0
# Add temporal derivative (EV 1)
set fmri(deriv_yn1) 0
# Custom EV file (EV 1)
set fmri(custom1) "/Desktop/FEAT_test/sub-MC002/ses-01/func/sub-MC002_ses-01_func_Occ_time_series.txt"
# Gamma sigma (EV 1)
set fmri(gammasigma1) 0
# Gamma delay (EV 1)
set fmri(gammadelay1) 0
# Orthogonalise EV 1 wrt EV 0
set fmri(ortho1.0) 0
# Orthogonalise EV 1 wrt EV 1
set fmri(ortho1.1) 0
# Contrast & F-tests mode
# real : control real EVs
# orig : control original EVs
set fmri(con_mode_old) orig
set fmri(con_mode) orig
# Display images for contrast_real 1
set fmri(conpic_real.1) 1
# Title for contrast_real 1
set fmri(conname_real.1) "Occ"
# Real contrast_real vector 1 element 1
set fmri(con_real1.1) 1
# Display images for contrast_orig 1
set fmri(conpic_orig.1) 1
# Title for contrast_orig 1
set fmri(conname_orig.1) "Occ"
# Real contrast_orig vector 1 element 1
set fmri(con_orig1.1) 1
# Contrast masking - use >0 instead of thresholding?
set fmri(conmask_zerothresh_yn) 0
# Do contrast masking at all?
set fmri(conmask1_1) 0
##########################################################
# Now options that don't appear in the GUI
# Alternative (to BETting) mask image
set fmri(alternative_mask) ""
# Initial structural space registration initialisation transform
set fmri(init_initial_highres) ""
# Structural space registration initialisation transform
set fmri(init_highres) ""
# Standard space registration initialisation transform
set fmri(init_standard) ""
# For full FEAT analysis: overwrite existing .feat output dir?
set fmri(overwrite_yn) 0
Screenshots / relevant information:
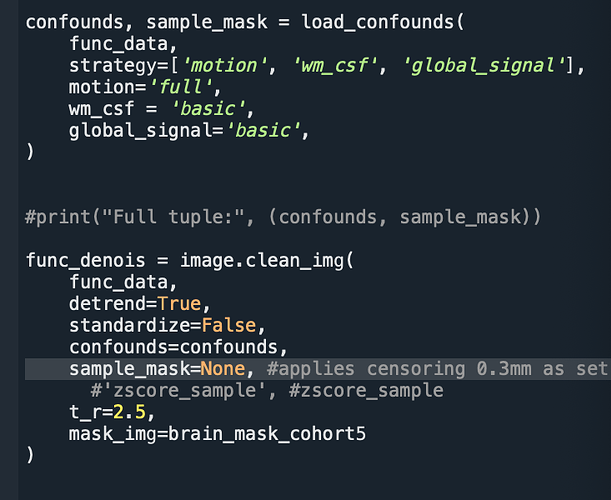
Confound set up and denoising code if helpful: