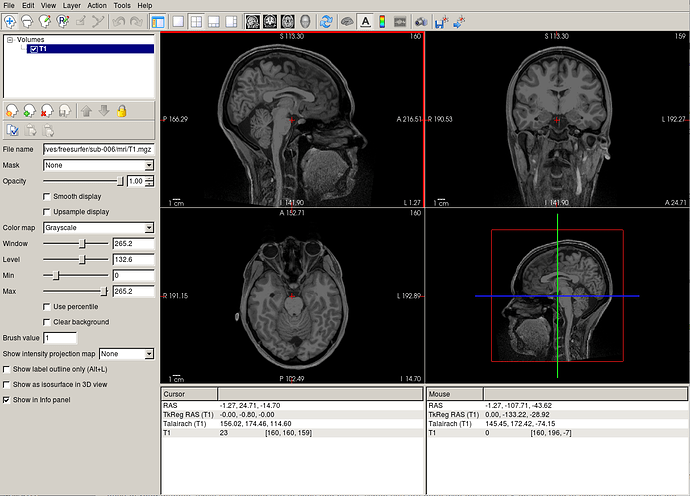
Hey, I am getting a weird error that I cannot find an answer for in about half of my subjects, showing one of them here as an example but all have similar errors:
File: /scratch/06873/zbretton/clearmem/derivatives/fmriprep/sub-006/log/20201012-105245_dcc56eac-6a15-46c6-953d-448b98d2a2d7/crash-20201012-120333-zbretton-autorecon2_vol-431a2155-a8cc-4125-8f56-49b7778b63f7.txt
Working Directory: /scratch/06873/zbretton/fmriprep/fmriprep_wf/single_subject_006_wf/anat_preproc_wf/surface_recon_wf/autorecon_resume_wf/autorecon2_vol
Inputs:
FLAIR_file:
T1_files:
T2_file:
args:
big_ventricles:
brainstem:
directive: autorecon2-volonly
environ: {}
expert:
flags:
hemi:
hippocampal_subfields_T1:
hippocampal_subfields_T2:
hires:
mprage:
mri_aparc2aseg:
mri_ca_label:
mri_ca_normalize:
mri_ca_register:
mri_edit_wm_with_aseg:
mri_em_register:
mri_fill:
mri_mask:
mri_normalize:
mri_pretess:
mri_remove_neck:
mri_segment:
mri_segstats:
mri_tessellate:
mri_watershed:
mris_anatomical_stats:
mris_ca_label:
mris_fix_topology:
mris_inflate:
mris_make_surfaces:
mris_register:
mris_smooth:
mris_sphere:
mris_surf2vol:
mrisp_paint:
openmp: 8
parallel:
steps:
subject_id: recon_all
subjects_dir:
talairach:
use_FLAIR:
use_T2:
xopts:
Traceback (most recent call last):
File "/usr/local/miniconda/lib/python3.7/site-packages/nipype/pipeline/plugins/multiproc.py", line 67, in run_node
result["result"] = node.run(updatehash=updatehash)
File "/usr/local/miniconda/lib/python3.7/site-packages/nipype/pipeline/engine/nodes.py", line 516, in run
result = self._run_interface(execute=True)
File "/usr/local/miniconda/lib/python3.7/site-packages/nipype/pipeline/engine/nodes.py", line 635, in _run_interface
return self._run_command(execute)
File "/usr/local/miniconda/lib/python3.7/site-packages/nipype/pipeline/engine/nodes.py", line 741, in _run_command
result = self._interface.run(cwd=outdir)
File "/usr/local/miniconda/lib/python3.7/site-packages/nipype/interfaces/base/core.py", line 419, in run
runtime = self._run_interface(runtime)
File "/usr/local/miniconda/lib/python3.7/site-packages/nipype/interfaces/base/core.py", line 814, in _run_interface
self.raise_exception(runtime)
File "/usr/local/miniconda/lib/python3.7/site-packages/nipype/interfaces/base/core.py", line 745, in raise_exception
).format(**runtime.dictcopy())
RuntimeError: Command:
recon-all -autorecon2-volonly -openmp 8 -subjid sub-006 -sd /scratch/06873/zbretton/clearmem/derivatives/freesurfer
Standard output:
Subject Stamp: freesurfer-Linux-centos6_x86_64-stable-pub-v6.0.1-f53a55a
Current Stamp: freesurfer-Linux-centos6_x86_64-stable-pub-v6.0.1-f53a55a
INFO: SUBJECTS_DIR is /scratch/06873/zbretton/clearmem/derivatives/freesurfer
Actual FREESURFER_HOME /opt/freesurfer
-rw-rw-r-- 1 zbretton G-814866 45022 Oct 12 11:58 /scratch/06873/zbretton/clearmem/derivatives/freesurfer/sub-006/scripts/recon-all.log
Linux nid00148 4.4.103-6.38_4.0.95-cray_ari_c #1 SMP Fri Feb 9 17:52:44 UTC 2018 (172b90b) x86_64 x86_64 x86_64 GNU/Linux
'/opt/freesurfer/bin/recon-all' -> '/scratch/06873/zbretton/clearmem/derivatives/freesurfer/sub-006/scripts/recon-all.local-copy'
#-------------------------------------
#@# EM Registration Mon Oct 12 12:02:46 CDT 2020
/scratch/06873/zbretton/clearmem/derivatives/freesurfer/sub-006/mri
mri_em_register -rusage /scratch/06873/zbretton/clearmem/derivatives/freesurfer/sub-006/touch/rusage.mri_em_register.dat -uns 3 -mask brainmask.mgz nu.mgz /opt/freesurfer/average/RB_all_2016-05-10.vc700.gca transforms/talairach.lta
setting unknown_nbr_spacing = 3
using MR volume brainmask.mgz to mask input volume...
== Number of threads available to mri_em_register for OpenMP = 8 ==
reading 1 input volumes...
logging results to talairach.log
reading '/opt/freesurfer/average/RB_all_2016-05-10.vc700.gca'...
average std = 7.3 using min determinant for regularization = 5.3
0 singular and 841 ill-conditioned covariance matrices regularized
reading 'nu.mgz'...
INFO: MRImask() using MRImaskDifferentGeometry()
INFO: MRImask() using MRImaskDifferentGeometry()
INFO: MRImask() using MRImaskDifferentGeometry()
INFO: MRImask() using MRImaskDifferentGeometry()
INFO: MRImask() using MRImaskDifferentGeometry()
freeing gibbs priors...done.
accounting for voxel sizes in initial transform
bounding unknown intensity as < 6.3 or > 503.7
total sample mean = 78.8 (1011 zeros)
************************************************
spacing=8, using 2830 sample points, tol=1.00e-05...
************************************************
register_mri: find_optimal_transform
find_optimal_transform: nsamples 2830, passno 0, spacing 8
GCAhistoScaleImageIntensities: could not find wm peak
resetting wm mean[0]: 98 --> 107
resetting gm mean[0]: 61 --> 61
input volume #1 is the most T1-like
using real data threshold=18.0
skull bounding box = (148, 26, 108) --> (171, 34, 131)
using (156, 29, 120) as brain centroid...
mean wm in atlas = 107, using box (153,28,117) --> (158, 29,122) to find MRI wm
before smoothing, mri peak at 105
robust fit to distribution - 73 +- 8.5
distribution too broad for accurate scaling - disabling
WARNING2: gca.c::GCAhistoScaleImageIntensities: h_mri->nbins=79, mri_peak=80
after smoothing, mri peak at 0, scaling input intensities by inf
Numerical result out of range
Linux nid00148 4.4.103-6.38_4.0.95-cray_ari_c #1 SMP Fri Feb 9 17:52:44 UTC 2018 (172b90b) x86_64 x86_64 x86_64 GNU/Linux
recon-all -s sub-006 exited with ERRORS at Mon Oct 12 12:03:32 CDT 2020
For more details, see the log file /scratch/06873/zbretton/clearmem/derivatives/freesurfer/sub-006/scripts/recon-all.log
To report a problem, see http://surfer.nmr.mgh.harvard.edu/fswiki/BugReporting
Standard error:
Return code: 1
Here is the end of the recon-all.log file:
#######################################
#-------------------------------------
#@# EM Registration Mon Oct 12 12:02:46 CDT 2020
/scratch/06873/zbretton/clearmem/derivatives/freesurfer/sub-006/mri
mri_em_register -rusage /scratch/06873/zbretton/clearmem/derivatives/freesurfer/sub-006/touch/rusage.mri_em_register.dat -uns 3 -mask brainmask.mgz nu.mgz /opt/freesurfer/average/RB_all_2016-05-10.vc700.gca transforms/talairach.lta
setting unknown_nbr_spacing = 3
using MR volume brainmask.mgz to mask input volume...
== Number of threads available to mri_em_register for OpenMP = 8 ==
reading 1 input volumes...
logging results to talairach.log
reading '/opt/freesurfer/average/RB_all_2016-05-10.vc700.gca'...
average std = 7.3 using min determinant for regularization = 5.3
0 singular and 841 ill-conditioned covariance matrices regularized
reading 'nu.mgz'...
INFO: MRImask() using MRImaskDifferentGeometry()
INFO: MRImask() using MRImaskDifferentGeometry()
INFO: MRImask() using MRImaskDifferentGeometry()
INFO: MRImask() using MRImaskDifferentGeometry()
INFO: MRImask() using MRImaskDifferentGeometry()
freeing gibbs priors...done.
accounting for voxel sizes in initial transform
bounding unknown intensity as < 6.3 or > 503.7
total sample mean = 78.8 (1011 zeros)
************************************************
spacing=8, using 2830 sample points, tol=1.00e-05...
************************************************
register_mri: find_optimal_transform
find_optimal_transform: nsamples 2830, passno 0, spacing 8
GCAhistoScaleImageIntensities: could not find wm peak
resetting wm mean[0]: 98 --> 107
resetting gm mean[0]: 61 --> 61
input volume #1 is the most T1-like
using real data threshold=18.0
skull bounding box = (148, 26, 108) --> (171, 34, 131)
using (156, 29, 120) as brain centroid...
mean wm in atlas = 107, using box (153,28,117) --> (158, 29,122) to find MRI wm
before smoothing, mri peak at 105
robust fit to distribution - 73 +- 8.5
distribution too broad for accurate scaling - disabling
WARNING2: gca.c::GCAhistoScaleImageIntensities: h_mri->nbins=79, mri_peak=80
after smoothing, mri peak at 0, scaling input intensities by inf
Numerical result out of range
Linux nid00148 4.4.103-6.38_4.0.95-cray_ari_c #1 SMP Fri Feb 9 17:52:44 UTC 2018 (172b90b) x86_64 x86_64 x86_64 GNU/Linux
recon-all -s sub-006 exited with ERRORS at Mon Oct 12 12:03:32 CDT 2020
To report a problem, see http://surfer.nmr.mgh.harvard.edu/fswiki/BugReporting
I have tried to search out for solutions but nothing seems to work. I have split up my subjects and tried to see if this was a memory issue, but that doesn’t seem the case. Running this in a singularity-docker, on TACC (Utexas HPC)
Any help is appreciated