Dear experts,
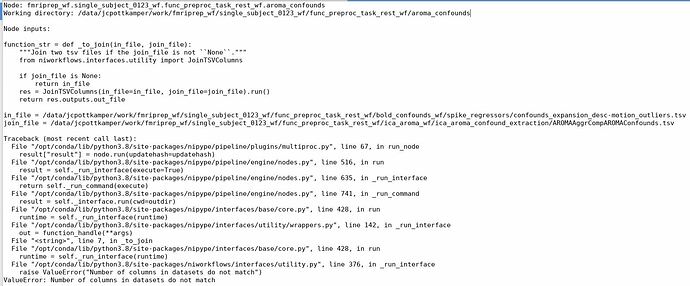
In our patient dataset, we do have a lot of movement and decided, based on the framewise displacement, to scratch the last volumes of excessive movement and run fmriprep again. It works up until a certain point. But we always get the error that fmriprep did not finish successfully (see picture) and it may have something to do with writing the confounds to the tsv file. We do get the AROMA nifti file, so in principle we can continue working with this output, but I find the error rather strange… Did anyone experience or can explain this? Thanks in advance.
Best,
Julia