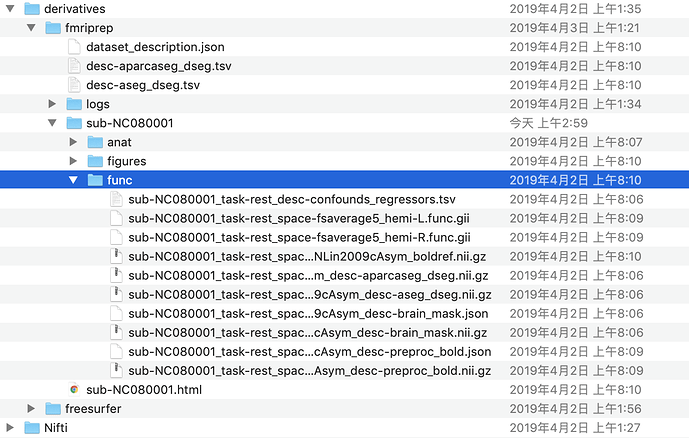
I’m getting this error now with 1-4-1rc5, but am assuming it’s pretty okay, except that I’m missing carpetplots. Sometimes it generates errors in the .html report, which are always in Node Name: fmriprep_wf.single_subject_NDARINV0CCEN5K2_wf.func_preproc_task_rest_run_[xx]_wf.carpetplot_wf.select_std, and sometimes not. An example of when it does:
[Node] Finished "fmriprep_wf.single_subject_NDARINV0CF1U8X8_wf.func_preproc_task_rest_run_01_wf.gen_cifti".
/usr/local/miniconda/lib/python3.7/site-packages/scipy/fftpack/basic.py:160: FutureWarning: Using a non-tuple sequence for multidimensional indexing is deprecated; use `arr[tuple(seq)]` instead of `arr[seq]`. In the future this will be interpreted as an array index, `arr[np.array(seq)]`, which will result either in an error or a different result.
z[index] = x
/usr/local/miniconda/lib/python3.7/site-packages/scipy/fftpack/basic.py:160: FutureWarning: Using a non-tuple sequence for multidimensional indexing is deprecated; use `arr[tuple(seq)]` instead of `arr[seq]`. In the future this will be interpreted as an array index, `arr[np.array(seq)]`, which will result either in an error or a different result.
z[index] = x
/usr/local/miniconda/lib/python3.7/site-packages/scipy/fftpack/basic.py:160: FutureWarning: Using a non-tuple sequence for multidimensional indexing is deprecated; use `arr[tuple(seq)]` instead of `arr[seq]`. In the future this will be interpreted as an array index, `arr[np.array(seq)]`, which will result either in an error or a different result.
z[index] = x
/usr/local/miniconda/lib/python3.7/site-packages/nipype/algorithms/confounds.py:1099: FutureWarning: `rcond` parameter will change to the default of machine precision times ``max(M, N)`` where M and N are the input matrix dimensions.
To use the future default and silence this warning we advise to pass `rcond=None`, to keep using the old, explicitly pass `rcond=-1`.
betas = np.linalg.lstsq(X, data.T)[0]
/usr/local/miniconda/lib/python3.7/site-packages/matplotlib/contour.py:1173: UserWarning: No contour levels were found within the data range.
warnings.warn("No contour levels were found"
/usr/local/miniconda/lib/python3.7/site-packages/nipype/algorithms/confounds.py:1099: FutureWarning: `rcond` parameter will change to the default of machine precision times ``max(M, N)`` where M and N are the input matrix dimensions.
To use the future default and silence this warning we advise to pass `rcond=None`, to keep using the old, explicitly pass `rcond=-1`.
betas = np.linalg.lstsq(X, data.T)[0]
/usr/local/miniconda/lib/python3.7/site-packages/nipype/algorithms/confounds.py:1099: FutureWarning: `rcond` parameter will change to the default of machine precision times ``max(M, N)`` where M and N are the input matrix dimensions.
To use the future default and silence this warning we advise to pass `rcond=None`, to keep using the old, explicitly pass `rcond=-1`.
betas = np.linalg.lstsq(X, data.T)[0]
/usr/local/miniconda/lib/python3.7/site-packages/matplotlib/contour.py:1173: UserWarning: No contour levels were found within the data range.
warnings.warn("No contour levels were found"
/usr/local/miniconda/lib/python3.7/site-packages/nipype/algorithms/confounds.py:1099: FutureWarning: `rcond` parameter will change to the default of machine precision times ``max(M, N)`` where M and N are the input matrix dimensions.
To use the future default and silence this warning we advise to pass `rcond=None`, to keep using the old, explicitly pass `rcond=-1`.
betas = np.linalg.lstsq(X, data.T)[0]
/usr/local/miniconda/lib/python3.7/site-packages/nipype/algorithms/confounds.py:1099: FutureWarning: `rcond` parameter will change to the default of machine precision times ``max(M, N)`` where M and N are the input matrix dimensions.
To use the future default and silence this warning we advise to pass `rcond=None`, to keep using the old, explicitly pass `rcond=-1`.
betas = np.linalg.lstsq(X, data.T)[0]
/usr/local/miniconda/lib/python3.7/site-packages/matplotlib/contour.py:1173: UserWarning: No contour levels were found within the data range.
warnings.warn("No contour levels were found"
/usr/local/miniconda/lib/python3.7/site-packages/nipype/algorithms/confounds.py:1099: FutureWarning: `rcond` parameter will change to the default of machine precision times ``max(M, N)`` where M and N are the input matrix dimensions.
To use the future default and silence this warning we advise to pass `rcond=None`, to keep using the old, explicitly pass `rcond=-1`.
betas = np.linalg.lstsq(X, data.T)[0]
Preprocessing did not finish successfully. Errors occurred while processing data from participants: NDARINV0CF1U8X8 (3). Check the HTML reports for details.
Error in atexit._run_exitfuncs:
Traceback (most recent call last):
File "/usr/local/miniconda/lib/python3.7/concurrent/futures/process.py", line 101, in _python_exit
thread_wakeup.wakeup()
File "/usr/local/miniconda/lib/python3.7/concurrent/futures/process.py", line 89, in wakeup
self._writer.send_bytes(b"")
File "/usr/local/miniconda/lib/python3.7/multiprocessing/connection.py", line 183, in send_bytes
self._check_closed()
File "/usr/local/miniconda/lib/python3.7/multiprocessing/connection.py", line 136, in _check_closed
raise OSError("handle is closed")
OSError: handle is closed
and when it doesn’t:
[Node] Finished "fmriprep_wf.single_subject_NDARINV0P4XZMZA_wf.func_preproc_task_rest_run_01_wf.bold_std_trans_wf.bold_reference_wf.enhance_and_skullstrip_bold_wf.apply_mask".
/usr/local/miniconda/lib/python3.7/site-packages/scipy/fftpack/basic.py:160: FutureWarning: Using a non-tuple sequence for multidimensional indexing is deprecated; use `arr[tuple(seq)]` instead of `arr[seq]`. In the future this will be interpreted as an array index, `arr[np.array(seq)]`, which will result either in an error or a different result.
z[index] = x
/usr/local/miniconda/lib/python3.7/site-packages/scipy/fftpack/basic.py:160: FutureWarning: Using a non-tuple sequence for multidimensional indexing is deprecated; use `arr[tuple(seq)]` instead of `arr[seq]`. In the future this will be interpreted as an array index, `arr[np.array(seq)]`, which will result either in an error or a different result.
z[index] = x
/usr/local/miniconda/lib/python3.7/site-packages/matplotlib/contour.py:1173: UserWarning: No contour levels were found within the data range.
warnings.warn("No contour levels were found"
/usr/local/miniconda/lib/python3.7/site-packages/nipype/algorithms/confounds.py:1099: FutureWarning: `rcond` parameter will change to the default of machine precision times ``max(M, N)`` where M and N are the input matrix dimensions.
To use the future default and silence this warning we advise to pass `rcond=None`, to keep using the old, explicitly pass `rcond=-1`.
betas = np.linalg.lstsq(X, data.T)[0]
/usr/local/miniconda/lib/python3.7/site-packages/scipy/fftpack/basic.py:160: FutureWarning: Using a non-tuple sequence for multidimensional indexing is deprecated; use `arr[tuple(seq)]` instead of `arr[seq]`. In the future this will be interpreted as an array index, `arr[np.array(seq)]`, which will result either in an error or a different result.
z[index] = x
/usr/local/miniconda/lib/python3.7/site-packages/nipype/algorithms/confounds.py:1099: FutureWarning: `rcond` parameter will change to the default of machine precision times ``max(M, N)`` where M and N are the input matrix dimensions.
To use the future default and silence this warning we advise to pass `rcond=None`, to keep using the old, explicitly pass `rcond=-1`.
betas = np.linalg.lstsq(X, data.T)[0]
/usr/local/miniconda/lib/python3.7/site-packages/matplotlib/contour.py:1173: UserWarning: No contour levels were found within the data range.
warnings.warn("No contour levels were found"
/usr/local/miniconda/lib/python3.7/site-packages/nipype/algorithms/confounds.py:1099: FutureWarning: `rcond` parameter will change to the default of machine precision times ``max(M, N)`` where M and N are the input matrix dimensions.
To use the future default and silence this warning we advise to pass `rcond=None`, to keep using the old, explicitly pass `rcond=-1`.
betas = np.linalg.lstsq(X, data.T)[0]
/usr/local/miniconda/lib/python3.7/site-packages/nipype/algorithms/confounds.py:1099: FutureWarning: `rcond` parameter will change to the default of machine precision times ``max(M, N)`` where M and N are the input matrix dimensions.
To use the future default and silence this warning we advise to pass `rcond=None`, to keep using the old, explicitly pass `rcond=-1`.
betas = np.linalg.lstsq(X, data.T)[0]
/usr/local/miniconda/lib/python3.7/site-packages/matplotlib/contour.py:1173: UserWarning: No contour levels were found within the data range.
warnings.warn("No contour levels were found"
/usr/local/miniconda/lib/python3.7/site-packages/scipy/fftpack/basic.py:160: FutureWarning: Using a non-tuple sequence for multidimensional indexing is deprecated; use `arr[tuple(seq)]` instead of `arr[seq]`. In the future this will be interpreted as an array index, `arr[np.array(seq)]`, which will result either in an error or a different result.
z[index] = x
/usr/local/miniconda/lib/python3.7/site-packages/nipype/algorithms/confounds.py:1099: FutureWarning: `rcond` parameter will change to the default of machine precision times ``max(M, N)`` where M and N are the input matrix dimensions.
To use the future default and silence this warning we advise to pass `rcond=None`, to keep using the old, explicitly pass `rcond=-1`.
betas = np.linalg.lstsq(X, data.T)[0]
/usr/local/miniconda/lib/python3.7/site-packages/nipype/algorithms/confounds.py:1099: FutureWarning: `rcond` parameter will change to the default of machine precision times ``max(M, N)`` where M and N are the input matrix dimensions.
To use the future default and silence this warning we advise to pass `rcond=None`, to keep using the old, explicitly pass `rcond=-1`.
betas = np.linalg.lstsq(X, data.T)[0]
/usr/local/miniconda/lib/python3.7/site-packages/nipype/algorithms/confounds.py:1099: FutureWarning: `rcond` parameter will change to the default of machine precision times ``max(M, N)`` where M and N are the input matrix dimensions.
To use the future default and silence this warning we advise to pass `rcond=None`, to keep using the old, explicitly pass `rcond=-1`.
betas = np.linalg.lstsq(X, data.T)[0]
/usr/local/miniconda/lib/python3.7/site-packages/nipype/algorithms/confounds.py:1099: FutureWarning: `rcond` parameter will change to the default of machine precision times ``max(M, N)`` where M and N are the input matrix dimensions.
To use the future default and silence this warning we advise to pass `rcond=None`, to keep using the old, explicitly pass `rcond=-1`.
betas = np.linalg.lstsq(X, data.T)[0]
/usr/local/miniconda/lib/python3.7/site-packages/matplotlib/contour.py:1173: UserWarning: No contour levels were found within the data range.
warnings.warn("No contour levels were found"
Error in atexit._run_exitfuncs:
Traceback (most recent call last):
File "/usr/local/miniconda/lib/python3.7/concurrent/futures/process.py", line 101, in _python_exit
thread_wakeup.wakeup()
File "/usr/local/miniconda/lib/python3.7/concurrent/futures/process.py", line 89, in wakeup
self._writer.send_bytes(b"")
File "/usr/local/miniconda/lib/python3.7/multiprocessing/connection.py", line 183, in send_bytes
self._check_closed()
File "/usr/local/miniconda/lib/python3.7/multiprocessing/connection.py", line 136, in _check_closed
raise OSError("handle is closed")
OSError: handle is closed
Preprocessing seems to finish fine for both, even in the second case where it looks like it stopped at rest_run_01_wf.bold_std_trans_wf.bold_reference_wf.enhance_and_skullstrip_bold_wf.apply_mask, but I don’t get any nice carpetplots in the .html file.
Edit: I just realized I reduced the amount of memory from 25 GBs down to 18-19 GBs…is this just a memory issue or something?