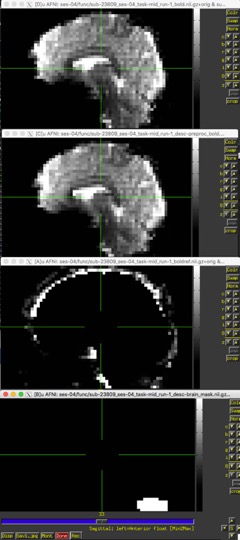
I’ve got a run with a brain mask that appears to be failing badly, as you can see in the image below. I’m not sure exactly where things are going wrong. Here’s a link to a zip file containing the subjects report:https://drive.google.com/open?id=1p_bcPRybbt8s0MBUYtW0O9eH-v2eMlo_
I’ve also got the work directory for that run saved which I can share if it will help diagnose the issue.
Ugh, even the least horrible masks are still horrible!
Does this dataset contain SBRefs? Could be an instance of https://github.com/poldracklab/fmriprep/issues/1684
Nope, no SBRefs, it does have field maps, but the orientation information on those matches the functional.
This may be caused by a failure of N4BiasFieldCorrection in cases where there are negative values in the input image. Issue posted to ANTS here: https://github.com/ANTsX/ANTs/issues/822
As a work around I’m going to try scaling the source functional so that it doesn’t have any negative values and see if that fixes the issue. Will update when that’s complete.
Alright, I take it back, there aren’t any negative values in the source data. The negative values are being introduced by interpolation in ANTS. See https://github.com/ANTsX/ANTs/issues/822. I’m not really sure what a temporary workaround for this would be, but workflows that use ANTS interpolation should maybe be drop negative values following interpolation.
Thanks a lot for your patience and the thorough testing - I’ve opened https://github.com/poldracklab/niworkflows/issues/388 to keep track of this problem.
Hi @Shotgunosine, we just released fMRIPrep 1.5.0 addressing these concerns. Could you check whether this issue has been fixed?
@oesteban I’ve run about 400 sessions through fmriprep under 1.4.1 and reviewed those results. Should the results from 1.5 be similar enough in most cases that I could use something like np.allclose() to check for differences or do I need to manually review all of those results again if I want to use 1.5.0?
I believe so. The biggest change between 1.4.1 and 1.5.0 is the split out of susceptibility distortion workflows to a new package. However, the code should be the same.
Other changes should not be impactfull, most of them related to singularity+templateflow+permissions.