Dear Chris,
I hope it’s ok to post this here, as we’ve encountered a very similar issue (using fmriprep v1.3.0.post3).
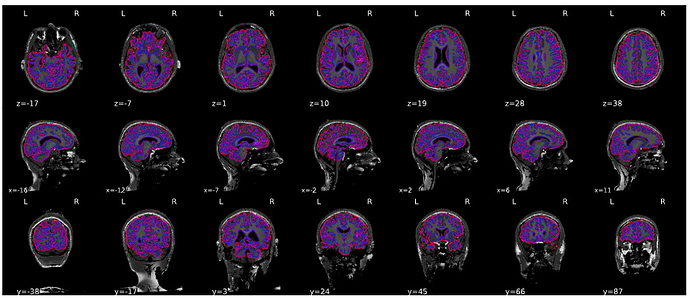
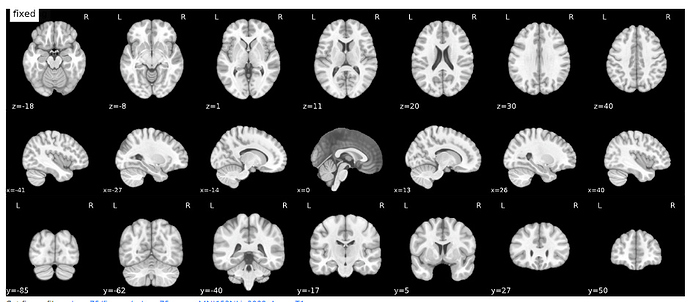
We see this problem in all of our volunteers though. I checked and the file you suggested indeed contains an over-cropped image.
It would be great if we could input our own brain mask (first we’d have to of course find an algorithm that works better for us, haven’t tried much yet except for bet). In the meantime, I wondered if you know of anything else that we could try?
Thanks,
Louise
1 Like
Hi Louise,
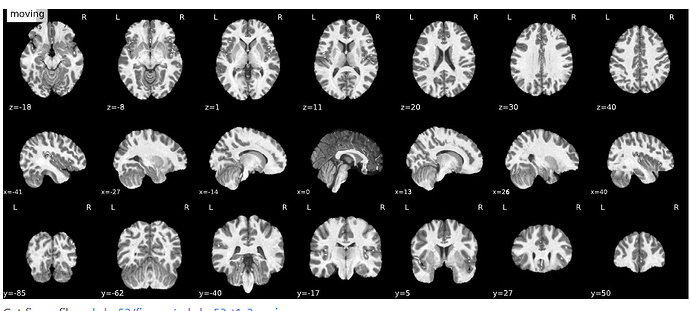
I’ve split the issue from the previous one because I’m convinced this is a different problem. In this case, the segmentation is aligned with the T1w. My wild guess is that bias field correction is not working very well for your case.
We have revised the brain extraction workflow and inhomogeneity correction. Could you give the pre-release we just posted a try?
It’s available via docker hub as poldracklab/fmriprep:1.4.0a0
Hi Oscar, thanks for your suggestion. I’ve got it running for a single subject right now, I will edit this message when it’s finished.
Update: seems like this fixed the issue! Thanks!
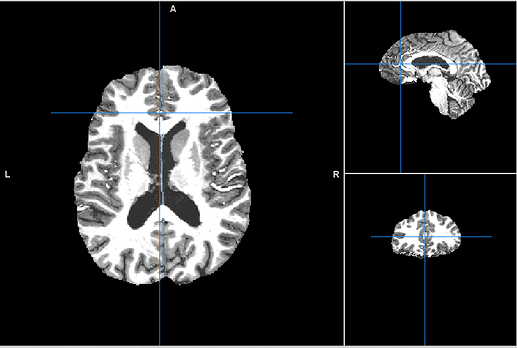
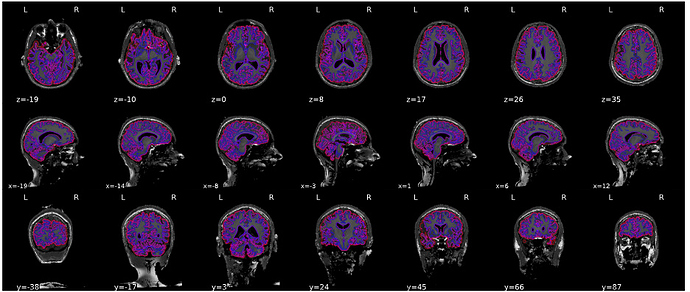
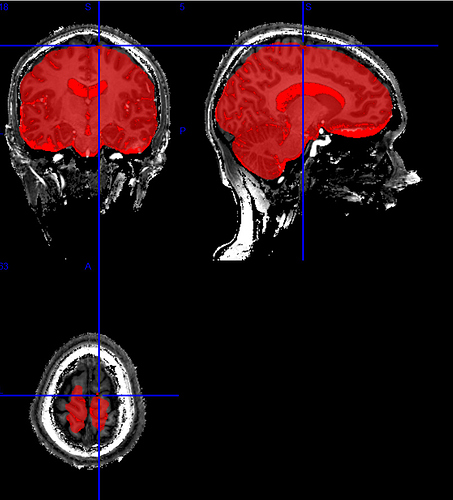
We had a closer look at the output and while the brain mask is much better than before, there still is some brain tissue not included in the mask. Here’s an example (created using fmriprep:1.4.0a0)
Should we continue with this, or is there maybe a way to fix this?
Thanks a lot for the feedback.
I think we are reaching the performance limitations of ANTs’ antsBrainExtraction workflow.
One possibility would be to change the target template. I’d be curious to see whether the results are better/worse using the MNI template in skull stripping. You’d need to use the --skull-strip-template MNI152NLin2009cAsym argument.
If that doesn’t work, then --skull-strip-template NKI could be a good last resort.
1 Like