hello stars
I have II ABIDE data arranged according to BIDS format.
Is there a command in the Freeserfer program that can be run to process all the data at once?
The number of images is more than 600, which is boring and very tiring to pass on each image separately to process it.
Thanks in advance
This depends on what your operating system is and what your memory and computing resources are like. Could you describe what you are working with?
Thank you very much for your cooperation.
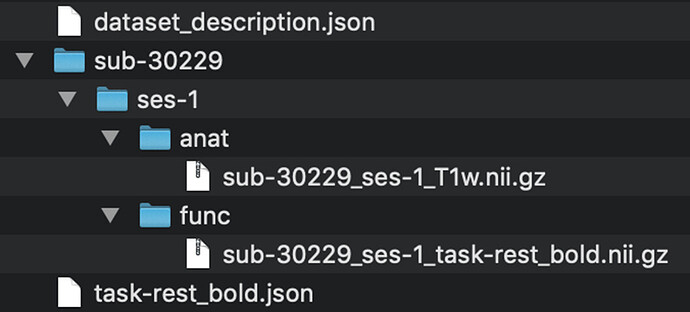
I have the data arranged as shown in the picture.
I am currently working on a virtual machine device after installing the Ubuntu system on it, there is 2 core for the vm . My host system it is Windows 10.
I also have access to a university supercomputer which is Linux. there are log node and computing node . Freesurfer is uploaded by the adminstors . And I don’t have permissions to download other programs .
In that case, your best bet is to use the supercomputer. This might help https://github.com/dfsp-spirit/shell-tools/tree/master/gnu_parallel_reconall_minimal.
Another way would be to do something like this
The submission script (submit_job_array.sh)
NOTE: Replace the all capitalized variables to match your needs.
#!/bin/bash
BIDS=PATH_TO_BIDS_DIR
pushd $BIDS
subjs=($(ls sub*/ -d)) # Get list of subject directory names
subjs=("${subjs[@]///}") # Remove the lagging slash
popd
# take the length of the array
# this will be useful for indexing later
len=$(expr ${#subjs[@]} - 1) # len - 1 because 0 index
echo Spawning ${#subjs[@]} sub-jobs. # print to command line
sbatch --array=0-$len single_subject_recon_all.sh $BIDS ${subjs[@]}
The script that defines the single subject recon-all command (single_subject_recon_all.sh)
Adjust the sbatch header as needed based on the requirements of your processing
#!/bin/bash
#SBATCH --time=1:00:00
#SBATCH --mem=8GB
#SBATCH --cpus-per-task=8
#SBATCH -J recon-all
args=($@)
BIDS=$1
subjs=(${args[@]:1})
sub=${subjs[${SLURM_ARRAY_TASK_ID}]}
echo $sub
input_niigz=$BIDS/$sub/ses-1/anat/${sub}_ses-1_T1w.nii.gz # Name of nifti input
gunzip $input_nii # Extract the nifti so it can be used in FreeSurfer
input_nii=${input_niigz//.gz} # File name without the .gz
recon-all -subjid $sub -i ${input_nii} # run recon-all
I haven’t tested this, but something like this should work and is how I parallelize jobs across my linux cluster.
How can I run this code after writing it. Sorry, this is my first interaction with Linux and I’m still learning
Just run ./submit_job_array.sh After putting the two files in the same directory.