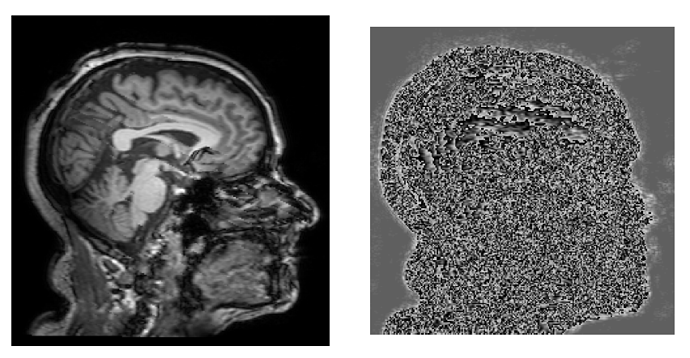
I have a T1w image which I am processing through an inhouse pipeline.
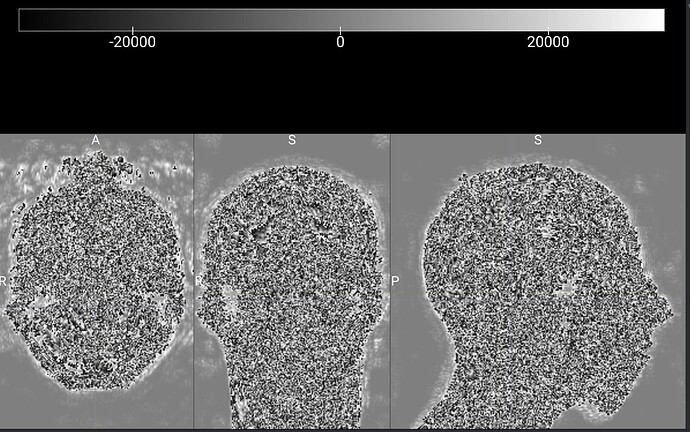
One of the steps, reorienting and bias field correction, causes a curious artifact
After digging into the pipeline, I found these steps were performed:
- Resample to LPI with
3dresample
- Reorient to MNI152 using
fslreorient2std
- Change filetype to nifti gz with
fslchfiletype
- N4 bias field correction using
N4BiasFieldCorrection
Replicating the steps, it appears that the 3rd step, with fslchfiletype, causes the image artifact. Checking the data types with fslinfo and mri_info does not reveal any change, although mri_info reports the image type changing from float to short.
I am curious, why is fslchfiletype obliterating the signal in this volume?
1 Like
First of all, cool artifact 
My guess: If you are changing the bytes that represent the values in the voxels but not update the nifti header correctly, the visualization software would interpret the bytes differently than what they actually mean. Therefore causing such artifacts.
I would recommend you to carefully track the data type of our voxels throughout the pipeline. fslinfo will just show you what is written in the header as the “intended” data type. Not exactly how the bytes are written in post header. Maybe try manipulating the data type header and see how the visualization software reacts.
Indeed, a weird one… I use it quite a lot that tool, and never faced a similar artifact.
If solution from @ofgulban doesnt work, consider using another package to change the format (such as mri_convert from freesurfer or ConvertImage from ANTs (since you already using N4correction…)
1 Like
What tool are you using to visualize the NIfTI images? Do the images look correct if you display open them with NiiVue, MRIcroGL or FSLeyes? The example you provide look like and endian error (e.g. the display software can only read big or little endian data).
1 Like
Images were viewed in freeview.
Viewing the image in fsleyes and MRIcroGL provide similar results
I will try out @ofgulban’s suggestion to see if that fixes anything.
1 Like
Changing filetype (via nib.Nifti1Image.set_data_dtype) does not alter the image shown in freeview, MRIcroGL, or when plotted via matplotlib.
The fsl commands above were ran using fsl version 6.0.0. Curiously, when running the same steps using fsl 6.0.5, the artifact is not reproduced…
See FSL release notes 6.0.2, 29/09/2019:
- Fixed ANALYZE support in fslchfiletype, plus fully report all known formats ( including NIFTI2 ).
I am going out on a limb here, but when you run fslhd on the image, what does it report as the data type. I am going to guess it is
data_type UINT16
Once upon a time, scanners used 12-bit ADCs, so magnitude images were in the range 0…4095. This fit nicely into the signed INT16 datatype that is native to FSL. Recently, scanners are using 16-bit ADC leading to magnitude voxels in the range of 0…65535. Old versions of FSL are unlikely to handle these. Modern versions of FSL (and AFNI) promote UINT16 to FLOAT32 internally.
1 Like
fslhd reports the image as follows:
image ran with fslchfiletype from FSL 6.0.0
data_type INT16
image ran with fslchfiletype from FSL 6.0.5.1
data_type FLOAT32
So, looks like your suspicions were correct.
1 Like
Closer inspection…although the data type is reported as INT16, the voxel values are actually stored as floats.
>>> import nibabel as nib
>>> a = nib.load('./problematic_t1.nii')
>>> b = a.get_fdata()
>>> c = b[b != 0]
>>> print(c)
array([1550.6751709, 1550.6751709, 1550.6751709, ..., 1550.6751709,
1550.6751709, 1550.6751709])
1 Like
Data stored as integers but multiplied by floating point scale slope and intercept.
Careful inspection of the colorbar in the MRIcroGL image also reveals the issue: notice the grayscale color bar range goes from -32768…+32767, yet by definition all voxels in a magnitude image are always positive. The reason is simple: magnitude is the vector length, and lengths are always positive regardless of direction. The magnitude is the Euclidean length of the real and imaginary component, which you can derive from the pythagorean theorem: length = sqrt(x^2 + y^2). Since any value multiplied by itself is positive, the pythagorean theorem always returns positive values.
2 Likes