Hey Neurostars,
I’ve used heudiconv before with little problem but I am running into one odd problem for one subject. There are 6 acquisitions of BOLD data.
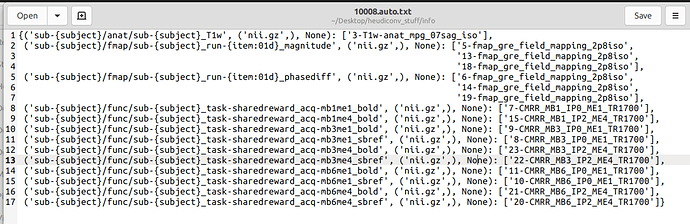
All of the acquisitions seem to be correctly identified when I look at the .auto.txt file in the .heudiconv directory
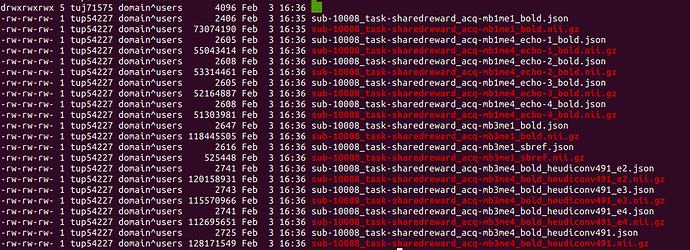
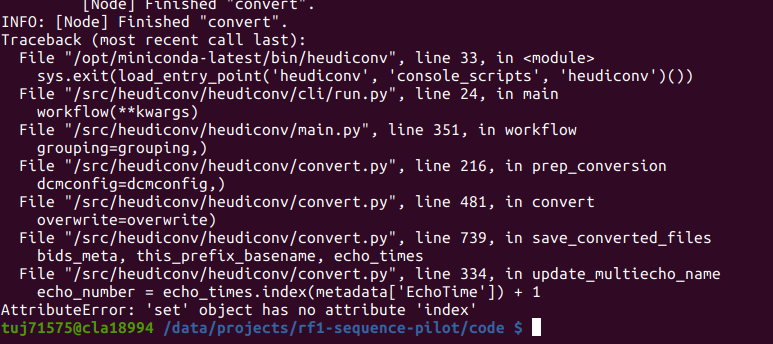
However, it adds an incorrect suffix to the mb3me4 acquisition and fails to convert the mb6me4 altogether.
It only seems to fail on the acquisitions that are both multiband and multi-echo however this isn’t a problem with another subject.
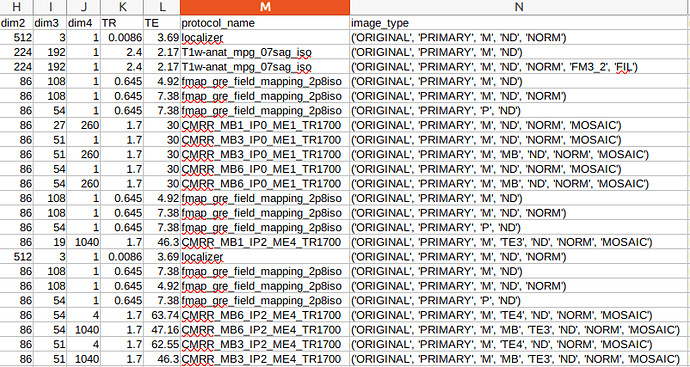
Here’s what the current heuristics file and dicominfo.tsv for the subject looks like. Any help would be appreciated
import os
def create_key(template, outtype=('nii.gz',), annotation_classes=None):
if template is None or not template:
raise ValueError('Template must be a valid format string')
return template, outtype, annotation_classes
def infotodict(seqinfo):
t1w = create_key('sub-{subject}/anat/sub-{subject}_T1w')
mag = create_key('sub-{subject}/fmap/sub-{subject}_run-{item:01d}_magnitude')
phase = create_key('sub-{subject}/fmap/sub-{subject}_run-{item:01d}_phasediff')
#me1
mb1me1 = create_key('sub-{subject}/func/sub-{subject}_task-sharedreward_acq-mb1me1_bold')
mb3me1 = create_key('sub-{subject}/func/sub-{subject}_task-sharedreward_acq-mb3me1_bold')
mb3me1_sbref = create_key('sub-{subject}/func/sub-{subject}_task-sharedreward_acq-mb3me1_sbref')
mb6me1 =create_key('sub-{subject}/func/sub-{subject}_task-sharedreward_acq-mb6me1_bold')
mb6me1_sbref = create_key('sub-{subject}/func/sub-{subject}_task-sharedreward_acq-mb6me1_sbref')
#me4
mb1me4 =create_key('sub-{subject}/func/sub-{subject}_task-sharedreward_acq-mb1me4_bold')
mb3me4 =create_key('sub-{subject}/func/sub-{subject}_task-sharedreward_acq-mb3me4_bold')
mb3me4_sbref =create_key('sub-{subject}/func/sub-{subject}_task-sharedreward_acq-mb3me4_sbref')
mb6me4 = create_key('sub-{subject}/func/sub-{subject}_task-sharedreward_acq-mb6me4_bold')
mb6me4_sbref = create_key('sub-{subject}/func/sub-{subject}_task-sharedreward_acq-mb6me4_sbref')
# mag: [],
# phase: [],
info = {t1w: [],mag: [],phase: [],
mb1me1: [],
mb3me1: [], mb3me1_sbref: [],
mb6me1: [], mb6me1_sbref: [],
mb1me4: [],
mb3me4: [], mb3me4_sbref: [],
mb6me4: [], mb6me4_sbref: [],}
list_of_ids = [s.series_id for s in seqinfo]
for s in seqinfo:
if ('T1w-anat_mpg_07sag_iso' in s.protocol_name) and ('NORM' in s.image_type):
info[t1w] = [s.series_id]
if ('gre_field' in s.protocol_name) and ('NORM' in s.image_type):
info[mag].append(s.series_id)
if ('gre_field' in s.protocol_name) and ('P' in s.image_type):
info[phase].append(s.series_id)
# no multi-echo
if (s.dim4 >= 150) and ('MB1_' in s.protocol_name) and ('_ME1' in s.protocol_name):
info[mb1me1].append(s.series_id)
idx = list_of_ids.index(s.series_id)
elif (s.dim4 >= 150) and ('MB3_' in s.protocol_name) and ('_ME1' in s.protocol_name):
info[mb3me1].append(s.series_id)
idx = list_of_ids.index(s.series_id)
info[mb3me1_sbref].append(list_of_ids[idx -1])
elif (s.dim4 >= 150) and ('MB6_' in s.protocol_name) and ('_ME1' in s.protocol_name):
info[mb6me1].append(s.series_id)
idx = list_of_ids.index(s.series_id)
info[mb6me1_sbref].append(list_of_ids[idx -1])
# multi-echo standard
if (s.dim4 >= 150) and ('MB1_' in s.protocol_name) and ('_ME4' in s.protocol_name):
info[mb1me4].append(s.series_id)
idx = list_of_ids.index(s.series_id)
elif (s.dim4 >= 150) and ('MB3_' in s.protocol_name) and ('_ME4' in s.protocol_name) :
info[mb3me4].append(s.series_id)
idx = list_of_ids.index(s.series_id)
info[mb3me4_sbref].append(list_of_ids[idx -1])
elif (s.dim4 >= 150) and ('MB6_' in s.protocol_name) and ('_ME4' in s.protocol_name):
info[mb6me4].append(s.series_id)
idx = list_of_ids.index(s.series_id)
info[mb6me4_sbref].append(list_of_ids[idx -1])
return info