Hi everyone,
I have a question about how to correctly orthogonalize filters in a standard fMRI preprocessing pipeline, especially in the context of temporal filtering and confound estimation.
The Lindquist et al. (2018) paper shows two approaches to do this. (1) Create an omnibus projection matrix consisting of all nuisance variables. (2) Orthogonalize a filter with respect to all previously removed sets of covariates to avoid re-introducing nuisance signals removed in previous steps. My question is about the second approach.
What do I want to do?
So, let’s say I want to perform the following preprocessing steps:
- Motion correction
- Slice timing correction
- Coregistration
- Temporal filtering
- Spatial filtering (i.e. smoothing)
Additionally, I want to compute the following confounds:
- DVARS (which takes a functional image as input)
- CompCor (which takes a functional image as input)
- Compute average signal in GM, WM, CSV, and whole brain
- Framewise displacement (takes estimated motion parameters)
- Compute 24 motion parameters according to Friston et. al. (takes estimated motion parameters)
My current approach
For the moment I ignore the effect of slice-time correction, coregistration or spatial filter. My goal is to orthogonalize the motion correction and temporal filters. If I understand the orthogonalization approach correctly, then it’s important that I also temporal filter the estimated motion parameters, before I correct for their influence.
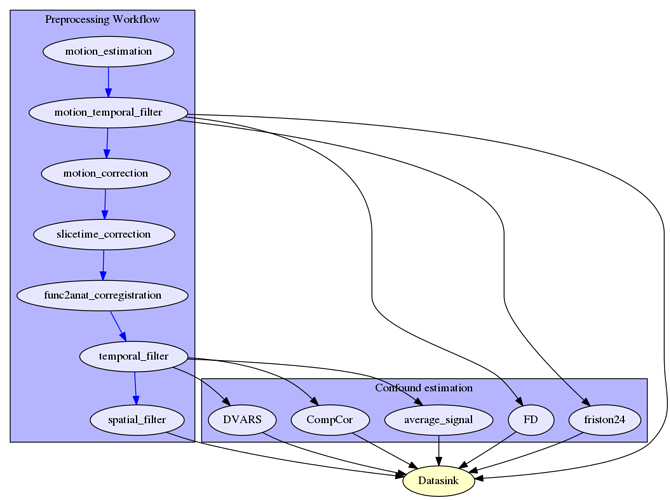
So, my approach would look something like this:
Questions
Does this seem right to you? I think the temporal filtering of the motion parameters is a good approach, especially in the context of high temporal acquisition, where the breathing pattern can be observed in the motion parameters.
But what about the confound estimations? Do I extract them correctly or would you do it differently?
Do I have to filter DVARS, CompCor, FD, Friston24 again after estimating?
Would you extract average_signal (and other confounds?) from the smoothed or unsmoothed image?
cheers,
michael