Hello Neurostars,
Though I originally used the output from mriqc this seemed to disagree with a metric that I was getting from MCFLIRT.
Both MRIQC ( “fd_mean” )& fmriprep (“rmsd” ) both cite that they implement the algorithm in Jenkinson, et al., 2002. also implemented in fsl’s MCFLIRT.
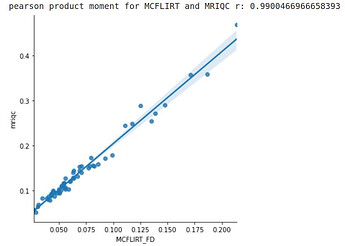
However, the MRIQC output was about 2 times as large as those from MCFLIRT and taking the mean of rmsd from fmriprep. I didn’t see anywhere in the MRIQC documentation where it explicitly states how it framewise displacement averaged or what units are output, but I assumed it took the mean in millimeters.
Any additional insight would be appreciated,
Best,
jsein
June 24, 2022, 9:18pm
2
My two cents
MRIQC can use AFNI 3dvolReg or FSL mcflirt whereas FMRIPREP uses only FSL mcflirt.
The methodology of the processing stages is also quite different between MRIQC and FMRIPREP due to the different goals of the two programs.
opened 08:12PM - 23 Jan 21 UTC
bug
### What version of fMRIPrep are you using?
FMRIPREP v 20.2.1
### What kin… d of installation are you using? Containers (Singularity, Docker), or "bare-metal"?
Singularity installation (version 2.6.1-dist), running on a high performance computing cluster.
### What is the exact command-line you used?
```
singularity run -B /scratch/jsein/BIDS:/work,$HOME/.templateflow:/opt/templateflow --cleanenv /scratch/jsein/my_images/fmriprep-20.2.1.simg \
--fs-license-file /work/freesurfer/license.txt /work/$study /work/$study/derivatives \
participant --participant-label $list_sub \
-w /work/temp_data_${study}\
--mem-mb 50000 --omp-nthreads 10 --nthreads 12 --ignore slicetiming --fd-spike-threshold 0.5 \
--output-spaces MNI152NLin2009cAsym MNI152NLin6Asym --return-all-components --cifti-output 91k
```
### Have you checked that your inputs are BIDS valid?
yes the input passes the bids-validator
### Did fMRIPrep generate the visual report for this particular subject? If yes, could you share it?
<!-- we can download links from Dropbox, Box, Google Drive, etc. You can send them privately to nipreps@gmail.com.
Reports do not contain data usable with personal identification or other research purposes -->
Yes the visual report was generated, no error reported, it is under the dropbox link at the end of this post.
### Can you find some traces of the error reported in the visual report (at the bottom) or in *crashfiles*?
no error reported
### Are you reusing previously computed results (e.g., FreeSurfer, Anatomical derivatives, work directory of previous run)?
No
### fMRIPrep log
If you have access to the output logged by fMRIPrep, please make sure to attach it as a text file to this issue.
We observed this strange output while building the GLM with nuisance regressors and noting that the columns with zeros where correlated with each other.
output: HTML reports, output and confounds_timeseries.tsv files under the following link:
https://www.dropbox.com/sh/3e455qbh8anqwl3/AAACn2-tnmRBhXx44M_vO0qEa?dl=0
Here is for example the content of the .par file output of mcflirt run by fmriprep: the zeros are very strange and it is a very different result from what I obtain if I use SPM for realigning the same run (but realignment is to the first scan instead to the sbref as it is set with fmriprep) No zeros are found with SPM.
EDIT: I just check with my local FSL installation and I found that mcflirt is giving the same output as fmriprep! Thus it is not seem to be a problem with fmriprep, rather a strange behaviour of mcflirt!
This subject had very low movements.
Thank you for sharing your thoughts!
```
0 -0 0 0 0 0
0 -0 0 0 0 0
0 -0 0 0 0 0
0 -0 0 0 0 0
0 -0 0 0 0 0.0298349
0 -0 0 0 0 0
0 -0 0 0 0 0
0 -0 0 0 0 0.0264596
0 -0 0 0 0 0.0125837
0 -0 0 0 0 0.00908859
0 -0 0 0 0 0.0499751
0 -0 0 0 0 0.0285046
0 -0 0 0 0 0.0394745
0 -0 0 0 0 0.032549
0 -0 0 0 0 0.0142394
0 -0 0 0 0 0.0188951
0 -0 0 0 0 0.0527396
0 -0 0 0 0 0.0118122
0 -0 0 0 0 0.0324686
-0.00020877 -0 0 0 0.00806986 0.0617947
-0.00071251 -0 0 0 0.0156193 0.0508352
0 -0 0 0 0 0.0259631
0 -0 0 0 0 0.0267916
0 -0 0 0 0 0.0423937
0 -0 0 0 0 0.0269359
0 -0 0 0 0 0.0480931
0 -0 0 0 0 0.0533892
-0.00079703 -0 0 0 0.0137821 0.0708697
-0.000779751 -0 0 0 0.011465 0.0502716
-0.00123065 -0 0 0 0.010698 0.109888
-0.00137164 -0 0 -7.29355e-09 0.0228613 0.0814645
-0.000797153 -0 0 -1.14295e-08 0.00402441 0.0433854
-0.00012957 -0 0 0 -0.000157725 0.0451129
-9.30172e-05 -0 0 0 -0.000177668 0.0566566
-0.000718193 -0 0 2.98428e-09 0.0150825 0.070767
-0.000450858 -0 0 9.52406e-09 0.00859951 0.0501525
-0.000182812 -0 0 0 0.00272729 0.0540445
-0.000355515 -0 0 0 0.00660644 0.056919
-0.00093645 -0 0 8.85655e-09 0.022218 0.0617246
-0.00130357 -0 0 0 0.0175132 0.0757637
-0.000789502 -0 0 0 0.0188825 0.0440254
-0.000264678 -0 0 0 0.00222485 0.0428122
-0.000721302 -0 0 0 0.0046294 0.0479397
-0.000891793 -0 0 8.90645e-09 0.021491 0.063626
-0.000721302 -0 0 0 0.0129002 0.0448448
-0.000112873 -0 0 0 -0.000384434 0.0460485
-0.000721302 -0 0 0 0.0136793 0.0609756
-0.000918056 -0 0 -4.79318e-09 0.0214994 0.0758632
-0.000987211 -0 0 1.42662e-08 0.0215346 0.0572181
-1.62193e-05 -0 0 1.0704e-09 -0.0050121 0.0533982
-0.000114733 -0 0 1.0704e-09 -0.000784969 0.0627668
-0.000738586 -0 0 1.0704e-09 0.0136945 0.0650518
-0.000980801 -0 0 9.64596e-09 0.0215313 0.0748123
-0.000738586 -0 0 1.0704e-09 0.00835188 0.0467987
-0.000738586 -0 0 1.0704e-09 0.00858199 0.0601027
-0.000738586 -0 0 1.0704e-09 0.00791957 0.027476
-0.000915425 -0 0 -3.87668e-09 0.0215165 0.0781238
-0.000632387 -0 0 1.07851e-08 0.010477 0.0423524
-0.00106174 -0 0 1.0704e-09 0.0250179 0.0660661
-0.000252066 -0 0 1.0704e-09 0.00448506 0.015156
-0.000550322 -0 0 1.0704e-09 0.00689665 0.0323721
-0.000738587 -0 0 1.0704e-09 0.0154435 0.0451829
-0.000738587 -0 0 1.0704e-09 0.0157021 0.0451829
-0.000683615 -0 0 8.63665e-09 0.00997629 0.0415626
-0.000738587 -0 0 1.0704e-09 0.0144991 0.0451829
-0.000738587 -0 0 1.0704e-09 0.0214398 0.062077
-0.000846131 -0 0 5.43473e-09 0.0214808 0.0616804
-0.000855447 -0 0 1.0704e-09 0.0142566 0.0805584
-0.000772026 -0 0 -1.47155e-09 0.021433 0.0404133
-0.000692231 -0 0 1.21844e-09 0.00489802 0.0434587
-0.000692231 -0 0 1.21844e-09 0.0135194 0.0451152
-0.000947193 -0 0 1.04547e-08 0.0275542 0.0769438
-0.000776082 -0 0 7.4812e-09 0.014442 0.0472611
-0.000674085 -0 0 2.25856e-09 0.00508311 0.0388478
-0.000674085 -0 0 2.25856e-09 0.0121347 0.0496988
-0.000787968 -0 0 -1.61307e-10 0.0214144 0.0552333
-0.000126402 -0 0 2.25856e-09 -0.00279745 0.0390678
-0.000793037 -0 0 7.2322e-09 0.021416 0.0833968
-0.000849837 -0 0 -8.19674e-09 0.0214422 0.0486355
-0.000752162 -0 0 2.25856e-09 0.0111124 0.0452097
-0.000752162 -0 0 2.99291e-09 0.0130734 0.0452097
-0.000962554 -0 0 2.99291e-09 0.0214808 0.0777942
-0.000142289 -0 0 2.99291e-09 0.0018386 0.0489621
-0.000752162 -0 0 2.99291e-09 0.0131606 0.0601034
-0.00089234 -0 0 -6.56433e-09 0.0214494 0.0828999
-0.000888647 -0 0 -6.16855e-09 0.0157029 0.0803104
-0.000844328 -0 0 -1.78585e-09 0.011968 0.0887311
-0.00169953 -0 0 -7.47431e-09 0.0334674 0.158842
-0.00087393 -0 0 5.37133e-09 0.0146743 0.0623504
0.000143044 -0 0 2.99291e-09 -0.00734077 0.0754622
-0.000691344 -0 0 -9.25198e-09 0.00984836 0.0752114
-0.000752162 -0 0 2.99291e-09 0.016751 0.084167
-0.000752163 -0 0 2.99291e-09 0.0157413 0.083586
-0.000752163 -0 0 2.99291e-09 0.00653493 0.0807244
-8.70316e-05 -0 0 2.99291e-09 0.00169775 0.0917994
-9.26849e-05 -0 0 2.99291e-09 0.00127143 0.0895586
-0.000752163 -0 0 2.99291e-09 0.017261 0.0870561
-0.00125371 -0 0 -1.00326e-08 0.0288015 0.14875
-0.000752163 -0 0 2.99291e-09 0.0146633 0.0639313
-0.000118441 -0 0 2.99291e-09 0.00127299 0.0817119
-0.000752163 -0 0 2.99291e-09 0.0129707 0.0823899
-0.000752163 -0 0 2.99291e-09 0.0213944 0.106449
-0.000752163 -0 0 2.99291e-09 0.0113494 0.0763368
-0.00065891 -0 0 -8.25317e-09 0.0111491 0.0822857
-0.000671937 -0 0 2.99291e-09 0.0112994 0.0823012
-0.000752163 -0 0 2.99291e-09 0.0128503 0.0925025
-0.000752163 -0 0 2.99291e-09 0.0155407 0.0901279
-0.000752163 -0 0 2.99291e-09 0.0168779 0.113742
-0.00089253 -0 0 7.76998e-09 0.0214438 0.0883661
-0.000679152 -0 0 1.24104e-08 0.00631072 0.0723094
-0.000657444 -0 0 -6.72726e-09 0.0106853 0.0822833
-0.000689144 -0 0 -3.30677e-09 0.0124616 0.0823191
-0.000925563 -0 0 7.72727e-09 0.0214587 0.108435
-0.000698344 -0 0 8.31619e-11 0.0157116 0.072331
-0.00068413 -0 0 -8.76388e-09 0.0117933 0.0863004
-0.000752164 -0 0 2.99291e-09 0.0161003 0.0883442
-0.000920858 -0 0 -5.52268e-09 0.0214595 0.113374
-0.000752164 -0 0 2.99291e-09 0.0147892 0.0723899
-0.000917793 -0 0 2.99291e-09 0.0214578 0.10781
-0.000689913 -0 0 2.29434e-10 0.0149052 0.0711225
-0.000566403 -0 0 9.45319e-09 0.00925237 0.0821727
-0.000752164 -0 0 2.99291e-09 0.0213944 0.0962025
-0.000752164 -0 0 2.99291e-09 0.0213944 0.0823899
-0.000638266 -0 0 1.16246e-08 0.0110292 0.0722563
-0.00066447 -0 0 2.99291e-09 0.0136406 0.0880102
-0.000752164 -0 0 2.99291e-09 0.0213944 0.0947306
-0.000651978 -0 0 -2.81807e-09 0.011058 0.0689206
-0.000667694 -0 0 1.15758e-08 0.0139397 0.0877071
-0.00154329 -0 0 2.99291e-09 0.0401142 0.136862
-0.00129329 -0 0 9.52764e-09 0.0260016 0.0759717
-0.000107472 -0 0 2.99291e-09 0.000271841 0.0816564
-2.47086e-05 -0 0 2.99291e-09 0.000230031 0.0877171
-5.01933e-05 -0 0 2.99291e-09 0.000241494 0.105102
-5.01933e-05 -0 0 2.99291e-09 0.000238228 0.0815856
-0.000111185 -0 0 2.99291e-09 0.000261973 0.0979274
-0.000300193 -0 0 2.99291e-09 0.00312824 0.113329
-0.000147117 -0 0 2.99291e-09 0.00027752 0.0988603
-5.01934e-05 -0 0 2.99291e-09 0.000244518 0.101356
-0.000482272 -0 0 2.99291e-09 0.0106453 0.139763
-5.01935e-05 -0 0 2.99291e-09 0.000247003 0.0815848
-0.000139604 -0 0 2.99291e-09 0.000282857 0.0957424
-0.000491681 -0 0 -1.30105e-08 0.0116485 0.119675
-5.01935e-05 -0 0 2.99291e-09 0.000251651 0.0815925
-7.66508e-05 -0 0 2.99291e-09 0.000259712 0.0977094
-5.01936e-05 -0 0 2.99291e-09 0.000249538 0.112486
-0.000558213 -0 0 2.99291e-09 0.0109131 0.112771
-5.01937e-05 -0 0 2.99291e-09 0.000246456 0.0977375
-0.000300194 -0 0 2.99291e-09 0.00514414 0.122534
-0.00134459 -0 0 1.54441e-08 0.0328532 0.141568
-0.000300194 -0 0 2.99291e-09 0.00815844 0.0883218
-0.000183202 -0 0 2.99291e-09 0.000293394 0.102353
-4.32239e-05 -0 0 2.99291e-09 0.000238674 0.107906
-0.00114743 -0 0 2.99291e-09 0.0290794 0.127425
-0.000897429 -0 0 1.00482e-08 0.0173483 0.0952883
-0.000889111 -0 0 1.04315e-08 0.0214945 0.105775
-0.00100408 -0 0 9.48222e-09 0.019849 0.11848
-0.00127335 -0 0 5.11865e-09 0.0311654 0.130845
-0.000296748 -0 0 2.08935e-09 0.00617523 0.0873417
-0.000969976 -0 0 1.85995e-09 0.0236238 0.109871
-0.00105306 -0 0 -8.34191e-10 0.0290198 0.101678
-0.000337284 -0 0 2.94065e-09 0.00976135 0.0935801
-0.000912522 -0 0 2.94065e-09 0.0217164 0.106609
-0.00100806 -0 0 1.09586e-08 0.0290003 0.0980294
-0.000235892 -0 0 2.94065e-09 0.00416129 0.0680161
-0.0002995 -0 0 -4.24463e-09 0.00763374 0.0718959
-0.000378003 -0 0 2.94065e-09 0.00766427 0.09784
-0.000476403 -0 0 1.142e-08 0.0105637 0.107273
-0.000264467 -0 0 1.70832e-08 0.00462611 0.0818563
-0.000378003 -0 0 2.94065e-09 0.00766427 0.107528
-0.00127529 -0 0 2.94065e-09 0.0279142 0.115822
-0.000394514 -0 0 2.94065e-09 0.0054216 0.0635719
-0.000338767 -0 0 2.94065e-09 0.00535792 0.0620108
-0.000310327 -0 0 2.94065e-09 0.00973431 0.0701747
-0.000510893 -0 0 -1.20349e-08 0.0121424 0.0721408
-0.000306212 -0 0 2.94065e-09 0.00654379 0.0608874
-0.000307715 -0 0 2.94065e-09 0.00973395 0.0745652
-0.000760893 -0 0 2.94065e-09 0.0166699 0.0886181
-0.000402248 -0 0 2.94065e-09 0.00976776 0.0587203
-0.00037274 -0 0 2.94065e-09 0.0097618 0.0704316
-0.00133981 -0 0 -7.1319e-09 0.0341621 0.118327
-0.00114327 -0 0 8.29037e-09 0.0243787 0.061372
-0.000397666 -0 0 1.7511e-09 0.00560246 0.048239
-0.000467144 -0 0 1.7511e-09 0.0135842 0.0720942
-0.000703884 -0 0 1.7511e-09 0.0136784 0.0925856
-0.000567805 -0 0 1.42559e-08 0.0136245 0.068412
-0.000456984 -0 0 1.7511e-09 0.0082714 0.082086
-0.00049742 -0 0 1.7511e-09 0.013601 0.0875239
-0.000579684 -0 0 6.21471e-09 0.0136322 0.0943948
-0.000886482 -0 0 -1.90109e-09 0.0178705 0.132457
-0.000392981 -0 0 1.7511e-09 0.00837947 0.0465562
-0.000453884 -0 0 1.7511e-09 0.00354411 0.0682447
-0.000455754 -0 0 1.7511e-09 0.00778869 0.076941
-0.00079791 -0 0 -2.25216e-09 0.0160182 0.119483
-0.000703884 -0 0 1.7511e-09 0.0136784 0.0860687
-0.000515806 -0 0 8.14077e-09 0.00505693 0.0721474
-0.000703884 -0 0 1.7511e-09 0.0136784 0.109038
-0.000973579 -0 0 1.7511e-09 0.0191724 0.0930738
-0.000703884 -0 0 1.7511e-09 0.00652437 0.102228
-0.00137622 -0 0 -3.02707e-09 0.0290181 0.121431
-0.000922695 -0 0 1.7511e-09 0.0164082 0.0597458
-0.000535682 -0 0 -1.52291e-08 0.00822229 0.0487303
-0.000416535 -0 0 1.7511e-09 0.0056854 0.0698911
-0.000675995 -0 0 1.52494e-08 0.0160388 0.101727
-0.000494376 -0 0 1.7511e-09 0.0135606 0.0598621
-0.000427305 -0 0 1.7511e-09 0.00713759 0.0711866
-0.00042642 -0 0 1.7511e-09 0.00950132 0.0675095
-0.000545192 -0 0 -1.2961e-08 0.0135841 0.0911589
-0.00113445 -0 0 4.96017e-09 0.0287818 0.0692903
-0.000248316 -0 0 1.7511e-09 0.0026122 0.0552848
-0.000296377 -0 0 1.7511e-09 0.00952237 0.0707775
-0.000984914 -0 0 -9.48523e-09 0.0220887 0.0928407
-0.00108519 -0 0 1.7511e-09 0.0242357 0.0728167
-0.000428385 -0 0 1.7511e-09 0.00972028 0.0664081
-0.000440844 -0 0 1.7511e-09 0.00669708 0.0700118
-0.000577341 -0 0 1.43718e-08 0.0147366 0.0949765
-0.000714186 -0 0 -8.42742e-09 0.0147911 0.0766545
-0.000567001 -0 0 1.7511e-09 0.00955145 0.0650231
-0.000952613 -0 0 1.7511e-09 0.0198898 0.107762
-0.000711122 -0 0 1.25251e-08 0.0147255 0.0502363
-0.000141859 -0 0 2.83079e-09 0.00133122 0.00681592
-0.000254764 -0 0 2.83079e-09 0.00438108 0.0245639
-0.000573799 -0 0 2.83079e-09 0.0144983 0.0500758
-0.000396892 -0 0 2.83079e-09 0.00894718 0.036291
-0.000450751 -0 0 1.14495e-08 0.0105671 0.0509599
-0.000477195 -0 0 -4.98444e-09 0.0144614 0.0619916
-0.000765275 -0 0 -2.22546e-09 0.0165784 0.104925
-0.000498349 -0 0 6.246e-09 0.00804579 0.0667286
-0.000573799 -0 0 2.83079e-09 0.00861331 0.107492
-0.000479679 -0 0 -4.89206e-09 0.00855949 0.0620016
-0.000323799 -0 0 2.83079e-09 0.00669057 0.0618326
-0.000505582 -0 0 1.70776e-08 0.0144763 0.0964717
-0.000483779 -0 0 -3.67766e-09 0.00838124 0.0620033
-0.000383789 -0 0 2.83079e-09 0.00331371 0.0780429
-0.000474073 -0 0 -5.8192e-10 0.00939916 0.0801106
-0.000573799 -0 0 2.83079e-09 0.0144983 0.0963335
-0.000475001 -0 0 -4.62251e-10 0.00791038 0.0719908
-0.000573799 -0 0 2.83079e-09 0.0144983 0.106045
-0.000484343 -0 0 -3.61993e-10 0.0144627 0.0481019
-0.000323799 -0 0 2.83079e-09 0.0056614 0.0513834
-0.00038702 -0 0 1.12477e-08 0.00775919 0.0662933
-0.000468504 -0 0 2.83079e-09 0.0144591 0.0814031
-0.000381009 -0 0 2.83079e-09 0.00618318 0.0502864
-0.000323799 -0 0 2.83079e-09 0.00439705 0.0645692
-0.000484721 -0 0 2.83079e-09 0.0144647 0.0776531
-0.000483708 -0 0 6.24865e-09 0.0144612 0.0556088
-0.000323799 -0 0 2.83079e-09 0.0102559 0.0534578
-0.000508544 -0 0 -8.55307e-09 0.0144758 0.0826264
-0.000475607 -0 0 2.83079e-09 0.0144603 0.055945
-0.00048232 -0 0 -8.61192e-10 0.0144635 0.0668524
-0.000573799 -0 0 2.83079e-09 0.0144983 0.055117
-0.000323799 -0 0 2.83079e-09 0.00819916 0.0483753
-0.000323799 -0 0 2.83079e-09 0.00439067 0.0618151
-0.000573799 -0 0 2.83079e-09 0.0144983 0.085265
-0.000477475 -0 0 2.83079e-09 0.0144557 0.045981
-0.000390461 -0 0 -6.07088e-09 0.00444682 0.0553409
-0.000477257 -0 0 6.35485e-09 0.0113832 0.0619901
-0.000750476 -0 0 1.25869e-08 0.0197093 0.107349
-0.000481706 -0 0 -3.52031e-10 0.0144592 0.0509564
-0.000323799 -0 0 2.83079e-09 0.0103062 0.055793
-0.00046289 -0 0 2.83079e-09 0.014456 0.0756933
-0.000495254 -0 0 8.29451e-09 0.0144655 0.0462053
-0.000373825 -0 0 2.83079e-09 0.0108125 0.051343
-0.000573799 -0 0 2.83079e-09 0.0144983 0.0721011
-0.000481413 -0 0 5.48884e-09 0.0144615 0.0403123
-0.000378721 -0 0 2.83079e-09 0.00595095 0.0527988
-0.000573799 -0 0 2.83079e-09 0.0144983 0.102255
-0.000400382 -0 0 -4.43367e-09 0.0069007 0.0287226
6.30042e-05 -0 0 2.83079e-09 -0.0026651 0.0452635
-0.000435495 -0 0 2.83079e-09 0.0097738 0.0547686
-0.000469106 -0 0 2.83079e-09 0.00982248 0.0560717
0.000125741 -0 0 2.83079e-09 -0.00412512 0.0484292
-0.000462934 -0 0 2.83079e-09 0.00751252 0.0650845
-0.000667804 -0 0 -5.24346e-09 0.0145207 0.0721987
-0.000424832 -0 0 3.74969e-09 0.00839867 0.0542887
-0.00050323 -0 0 3.74969e-09 0.0144516 0.083276
-0.00118624 -0 0 3.74969e-09 0.0289049 0.10322
-0.000578415 -0 0 3.74969e-09 0.0157659 0.0492656
-0.000412126 -0 0 3.74969e-09 0.00332497 0.054508
-0.000404368 -0 0 3.74969e-09 0.0123336 0.0578388
-0.00051215 -0 0 9.10393e-09 0.0123738 0.0910823
-0.000404368 -0 0 3.74969e-09 0.00882121 0.0543322
-0.000404368 -0 0 3.74969e-09 0.0123336 0.0539334
-0.000404368 -0 0 3.74969e-09 0.00871471 0.0561
-0.000404368 -0 0 3.74969e-09 0.00873877 0.0555488
-0.000507047 -0 0 -5.76854e-09 0.0123691 0.0897949
-0.000768395 -0 0 1.24991e-08 0.0158443 0.0859467
-0.000392008 -0 0 2.32531e-09 0.00623554 0.0343081
-0.000392008 -0 0 2.32531e-09 0.0122711 0.0456019
-0.000392008 -0 0 2.32531e-09 0.0122711 0.0526101
-0.000392008 -0 0 2.32531e-09 0.00931002 0.0517353
-0.000392008 -0 0 2.32531e-09 0.00584972 0.0423322
-0.000392008 -0 0 2.32531e-09 0.0122711 0.0527413
-0.000392008 -0 0 2.32531e-09 0.0122711 0.0619527
-0.000537575 -0 0 -2.55388e-09 0.012324 0.0721128
-0.000457688 -0 0 -3.89321e-10 0.0122959 0.0840514
-0.000474748 -0 0 -6.42473e-10 0.0123049 0.0461615
-0.000392008 -0 0 2.32531e-09 0.00542904 0.0464291
-0.000392008 -0 0 2.32531e-09 0.0122711 0.0504269
-0.000392008 -0 0 2.32531e-09 0.0122711 0.0719527
-0.000392008 -0 0 2.32531e-09 0.0122711 0.0507589
-0.000392008 -0 0 2.32531e-09 0.00858723 0.0541215
-0.000392008 -0 0 2.32531e-09 0.008138 0.0656587
-0.000551162 -0 0 2.32531e-09 0.0123354 0.0721295
-0.000392008 -0 0 2.32531e-09 0.00491105 0.0497302
-0.000482046 -0 0 -6.23686e-10 0.0123049 0.0720546
-0.0006911 -0 0 2.32531e-09 0.0194784 0.0669943
-0.000360711 -0 0 2.32531e-09 0.00555064 0.0250721
-0.000347486 -0 0 2.32531e-09 0.00468455 0.0332049
-0.000388303 -0 0 2.32531e-09 0.0122163 0.0419559
0.000231651 -0 0 2.32531e-09 -0.00607035 0.0548363
-0.000304605 -0 0 2.32531e-09 0.0079979 0.0580075
-0.000170488 -0 0 2.32531e-09 0.0029905 0.0312546
0.000271837 -0 0 -1.80617e-09 -0.00468443 0.0380728
0.000268027 -0 0 1.17999e-08 -0.00618964 0.0445663
0.000261084 -0 0 1.35479e-08 -0.00598816 0.0452548
0.000181061 -0 0 2.32531e-09 -0.00862791 0.0828628
0.00036988 -0 0 2.32531e-09 -0.0119828 0.0501614
0.00027085 -0 0 1.25974e-08 -0.00556844 0.0679252
0.000181061 -0 0 2.32531e-09 -0.0060663 0.0913342
0.000276382 -0 0 -2.61899e-09 -0.0119468 0.0343135
0.000274122 -0 0 2.32531e-09 -0.00934721 0.0487565
0.000322834 -0 0 1.18787e-08 -0.0061962 0.0650543
0.000181061 -0 0 2.32531e-09 -0.00699194 0.0828496
0.000181061 -0 0 2.32531e-09 -0.00716909 0.0417544
0.000339541 -0 0 -5.74701e-09 -0.00958281 0.0612099
-0.000146911 -0 0 2.32531e-09 0.00494863 0.105116
0.000181061 -0 0 2.32531e-09 -0.00365363 0.0208984
0.000288511 -0 0 2.32531e-09 -0.0119517 0.0223716
0.00027053 -0 0 6.85316e-09 -0.00629294 0.0500681
-0.000236648 -0 0 2.32531e-09 0.00532296 0.0606858
0.000181061 -0 0 2.32531e-09 -0.00860397 0.0279844
0.000244789 -0 0 -2.63327e-09 -0.00729157 0.0500972
2.38631e-06 -0 0 2.32531e-09 0.00281907 0.0794415
0.000181061 -0 0 2.32531e-09 -0.00350589 0.0501742
0.000181061 -0 0 2.32531e-09 -0.00955198 0.0625052
-2.97965e-06 -0 0 2.32531e-09 0.00210593 0.0949698
0.000181061 -0 0 2.32531e-09 -0.00830979 0.043568
0.000298899 -0 0 -5.45857e-09 -0.0119505 0.0534515
0.000181061 -0 0 2.32531e-09 -0.00190411 0.0874287
0.000181061 -0 0 2.32531e-09 -0.00374029 0.0634447
0.000317519 -0 0 2.32531e-09 -0.0119585 0.0529064
0.000181061 -0 0 2.32531e-09 -0.00312477 0.0772102
7.06293e-05 -0 0 2.32531e-09 0.00037782 0.0733328
0.000289854 -0 0 7.07189e-09 -0.00840966 0.0473801
0.000331191 -0 0 -5.55191e-09 -0.0119579 0.0599957
-0.000241041 -0 0 2.32531e-09 0.0050688 0.0806819
0.000181061 -0 0 2.32531e-09 -0.00190411 0.0167831
0.000181061 -0 0 2.32531e-09 -0.00844981 0.0195735
0.000340188 -0 0 2.32531e-09 -0.0119715 0.0361546
0.000181061 -0 0 2.32531e-09 -0.00597024 0.031775
0.000181061 -0 0 2.32531e-09 -0.00214056 0.0480441
0.000181061 -0 0 2.32531e-09 -0.00407231 0.015303
0.000181061 -0 0 2.32531e-09 -0.00520885 0.0195735
0 -0 0 0 0 0.0343214
```
Here is a plot (made with FSLeyes) of the translation along x, calculated with mcflirt (.par file, 4th column, ref= sbref) and with spm (rp file, 1st column, ref=1st scan): Quite different!

Another example: same run, looking at the ortation around y, calculated with mcflirt (.par file, 2nd column, ref= sbref) and with spm (rp file, 5th column, ref=1st scan):

jsein
June 24, 2022, 9:28pm
3
Regarding your experiment, how did you choose your theshold in FD? I was wondering was would be the relationship of the FD threshold and the length of the TR for instance.
Also one technical note: The coil used may have an influence as coil with high receive heterogeneity may overestimate the motion parameters: see here:
Especially in this paragraph:
What if we can’t even attain the physiologic noise-limiting regime? It’s quite possible to be in a subject motion-limiting regime, as anyone who has run an fMRI experiment can attest. In that case, the use of a high dimensional array coil (of 32 channels, say) could actually impose a higher motion sensitivity on the time series than it would have had were it detected by a smaller array coil (of 12 channels, say), due to the greater receive field heterogeneity of the 32-channel coil. This was something a colleague and I considered last year, in an arXiv paper ([1210.3633] A Simulation of the Effects of Receive Field Contrast on Motion-Corrected EPI Time Series ) and accompanying blog post . In an e-poster at this year’s ISMRM Annual Meeting (abstract #3352 ; a PDF of the slides is available via this Dropbox link ) we simulated the effects of motion on temporal SNR (tSNR), as well as the potential for spurious correlations in resting-state fMRI, when using a 32-channel coil. In doing these simulations we assumed perfect motion correction yet there were still drastic effects, as the above figure illustrates.
Hey @jsein ,
Sorry, I was actually I was mistaken about the threshold we chose. We were looking at excluding runs with mean framewise displacement greater than 0.5mm or greater than 1.35 (in our case about 1/2 a voxel width) in absolute head motion from the initial time point. Actually, despite the fact that the measure of mean framewise displacement from mriqc is about 2 times what comes out of fmriprep and mcflirt none are greater than 0.5 so this is more of a learning experience for me.
In our experiment the TR actually varies a bit across some acquisitions that we’re testing through but we have the same exclusion criteria for each run/acquisition.
We are using a headcoil with 64 channels at the moment but had done some piloting with a lower number. I can take a quick look and compare those sets as well to see if that’s a reasonable source of this problem.