Hi all!
I’m currently working with a data-set on diffusion MRI data and am trying to do a connectivity-based classification using k-means. From the FSL user guide, I found the following script to run the algorithm in MATLAB:
% Load Matrix2
x=load(‘fdt_matrix2.dot’);
M=full(spconvert(x));
% Calculate cross-correlation
CC = 1+corrcoef(M’);
% Do kmeans with k clusters
idx = kmeans(CC,k); % k is the number of clusters
% Load coordinate information to save results
addpath([getenv(‘FSLDIR’) ‘/etc/matlab’]);
[mask,~,scales] = read_avw(‘fdt_paths’);
mask = 0*mask;
coord = load(‘coords_for_fdt_matrix2’)+1;
ind = sub2ind(size(mask),coord(:,1),coord(:,2),coord(:,3));
[~,~,j] = unique(idx);
mask(ind) = j;
save_avw(mask,‘clusters’,‘i’,scales);
!fslcpgeom fdt_paths clusters
I would like to run this clustering algorithm for the one brain cluster (left and right sides) for each subject.
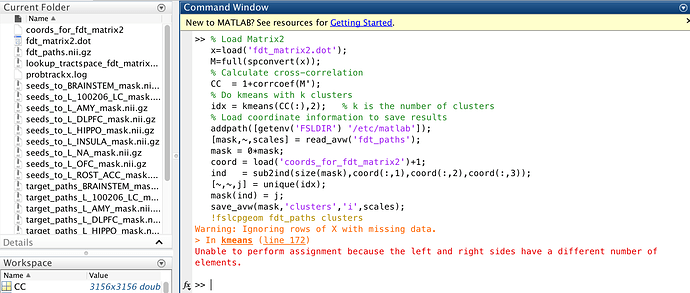
I’ve replaced idx = kmeans (CC(:),2); % k is the number of clusters and am getting the following error when running the script:
Warning: Ignoring rows of X with missing data.
In kmeans (line 172)
Unable to perform assignment because the left and right sides have different number of elements.
I am very new to MATLAB and unfortunately am not understanding how to quite get around this problem. I would greatly appreciate any help you can provide with this. Thank you so much in advance!
I’m also including a screenshot and leaving the link to script below: