Hi everyone,
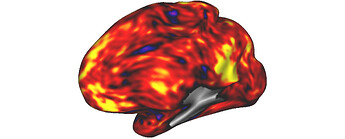
I’m new to analysing Human Connectome Project Young Adult (HCP-YA) data, and I’m finding the surface-based processing a bit challenging. As a test project, I aimed to create a functional map of the Default Mode Network (DMN) for one participant. I used the activity in the Posterior Cingulate Cortex (PCC) as a reference and calculated its correlation with the entire brain. Using nilearn and the Connectome Workbench, I successfully generated a map that meets my expectations when viewed in the participant’s native surface mesh:
However, I run into problems when I try to transfer this map to a standard fsaverage_LR32k surface. The map appears distorted in this standard space, and I’m not sure why. Here is a visualisation of the results using the surface found in T1w/MNINonLinear/fsaverage_LR32k:
Here is the code that I used to move the correlation map from one surface to the other:
wb_command -metric-resample \
/100206.rfMRI_REST1_LR.R.native.PCC_seed_corr_smoothed.func.gii \
/T1w/MNINonLinear/Native/100206.R.sphere.native.surf.gii \
/T1w/MNINonLinear/fsaverage_LR32k/100206.R.sphere.32k_fs_LR.surf.gii \
ADAP_BARY_AREA \
/100206.rfMRI_REST1_LR.R.32k_fs_LR.PCC_seed_corr_smoothed.func.gii \
-area-surfs \
/T1w/MNINonLinear/Native/100206.R.midthickness.native.surf.gii \ /T1w/MNINonLinear/fsaverage_LR32k/100206.R.midthickness.32k_fs_LR.surf.gii
I would greatly appreciate any guidance on what I might be doing incorrectly or how I could troubleshoot this issue. Thank you!