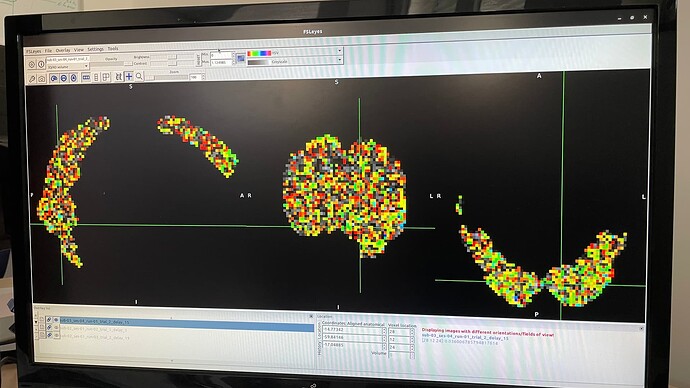
I want to extract maps specific to each Trial and each FIR delay of a run as input for my decoding analysis.
For some trials, the contrast is computed, but results in a map that is missing a lot of information in the brain. As there is no error message and it does not happen all of the time but just in some cases, I cannot figure out what is going wrong. In the “empty” spaces, the map contains of zeros.
Hi @zieranna and welcome to neurostars!
It will be hard for us to help without seeing a code example used to create these images. In particular it would be valuable to see how a brain mask is being defined.
Best,
Steven
My code is:
fir = FirstLevelModel(t_r=1.2, hrf_model="fir", fir_delays=[1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12,
13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25], standardize = False, signal_scaling=False)
for r in range(0, len(fmri_img), 2):
fir.fit([fmri_img[r],[[fmri_img[r+1]], events= [event_glm[r], event_glm[r+1]])
design_matrix = pd.DataFrame(fir.design_matrices[0])
design_matrix2 = pd.DataFrame(fir.design_matrices[1])
column_all = np.unique((design_matrix.columns, design_matrix2.columns))
contrasts = {
column: np.array((np.zeros(len(design_matrix.columns)),
np.zeros(len(design_matrix2.columns))))
for i, column in enumerate(column_all) }
for m, mat in enumerate(contrasts.keys()):
for c1, col1 in enumerate(design_matrix.columns):
if col1 == mat:
contrasts[str(mat)][0][c1] = 1
for c2, col2 in enumerate(design_matrix2.columns):
if col2==mat:
contrasts[str(mat)][1][c1] = 1
for i, column in tqdm(enumerate(contrasts.keys()):
t_map = fir.compute_contrast((contrasts[column][0], contrasts[column][1]), stat_type='t', output_type='stat')
t_map.to_filename()
I am always fitting two runs, because my conditions are randomized across the two of them.
Do you have a brain mask that you can pass in with the FirstLevelModel argument mask_img?