Attached is the result of crashfiles;
File: /media/nufas/ba32d503-d5ae-47f8-94a6-5318a1ce1fdf/home/farahana/Documents/dataset/crash-20180308-203349-nufas-selectfiles.a0-fcdd7b0c-9a49-4d9c-9638-9223ed367e8a.pklz
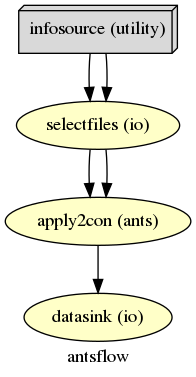
Node: antsflow.selectfiles
Working directory: /media/nufas/ba32d503-d5ae-47f8-94a6-5318a1ce1fdf/home/farahana/Documents/dataset/ds109_raw/output/workingdir/antsflow/_fwhm_id_4_subject_id_sub005/selectfiles
Node inputs:
base_directory = ds109_raw/output/
force_lists = False
fwhm_id = 4
ignore_exception = False
raise_on_empty = True
sort_filelist = True
subject_id = sub005
Traceback:
Traceback (most recent call last):
File "/home/nufas/anaconda3/lib/python3.6/site-packages/nipype/pipeline/plugins/linear.py", line 44, in run
node.run(updatehash=updatehash)
File "/home/nufas/anaconda3/lib/python3.6/site-packages/nipype/pipeline/engine/nodes.py", line 487, in run
result = self._run_interface(execute=True)
File "/home/nufas/anaconda3/lib/python3.6/site-packages/nipype/pipeline/engine/nodes.py", line 571, in _run_interface
return self._run_command(execute)
File "/home/nufas/anaconda3/lib/python3.6/site-packages/nipype/pipeline/engine/nodes.py", line 650, in _run_command
result = self._interface.run(cwd=outdir)
File "/home/nufas/anaconda3/lib/python3.6/site-packages/nipype/interfaces/base/core.py", line 517, in run
outputs = self.aggregate_outputs(runtime)
File "/home/nufas/anaconda3/lib/python3.6/site-packages/nipype/interfaces/base/core.py", line 591, in aggregate_outputs
predicted_outputs = self._list_outputs()
File "/home/nufas/anaconda3/lib/python3.6/site-packages/nipype/interfaces/io.py", line 1406, in _list_outputs
raise IOError(msg)
OSError: No files were found matching con template: /media/nufas/ba32d503-d5ae-47f8-94a6-5318a1ce1fdf/home/farahana/Documents/dataset/ds109_raw/output/workingdir/antsflow/_fwhm_id_4_subject_id_sub005/selectfiles/ds109_raw/output/datasink/1stLevel/sub005/fwhm-4/???_00??.nii