I’ve probably finished downloading the qsiprep-1.0.0.sif image, and I’ve replaced it with other available image sources with the following commands:
singularity build qsiprep-1.0.0.sif docker://swr.cn-north-4.myhuaweicloud.com/ddn-k8s/docker.io/pennlinc/qsiprep:latest
The following are the (partial) results of the image download after this command is executed:
===============================================================] 100 % 14.7 MiB/s 0s
155.8MiB / 155.8MiB [=============================================================================================================================================================================] 100 % 14.7 MiB/s 0s
111.2MiB / 111.2MiB [=============================================================================================================================================================================] 100 % 14.7 MiB/s 0s
87.3MiB / 87.3MiB [===============================================================================================================================================================================] 100 % 14.7 MiB/s 0s
199.1MiB / 199.1MiB [=============================================================================================================================================================================] 100 % 14.7 MiB/s 0s
INFO: Extracting OCI image...
INFO: Inserting Singularity configuration...
INFO: Creating SIF file...
INFO: Build complete: qsiprep-1.0.0.sif
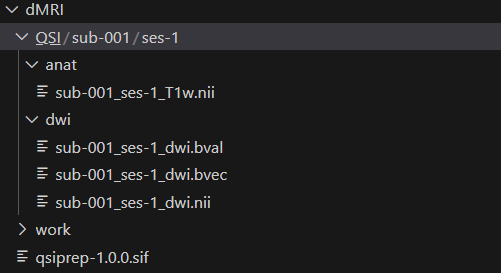
After acquiring the image, I would like to test if QSIPrep is available using some prepared test data. The data I prepared is shown below with the root directory dMRI and the root directory for the BIDs data QSI.
I tried to define my own slurm script by referring to the posts of experienced QSIPrep users within the forum (QSIPrep Optimal Compute Resources):
#!/bin/bash
#SBATCH -J QSI
#SBATCH -p bme_cpu
#SBATCH --mail-type=fail
#SBATCH -o QSI.out
#SBATCH -e QSI.err
#SBATCH --mem=50G
#SBATCH -N 1
#SBATCH --time=24:00:00
module load /public/software/modules/tools/singularity/3.5.2
#module load /public/software/modules/tools/singularity/4.1.4
bids_dir=/public_bme2/bme-zhanghan/Jiayifan/dMRI/QSI
participant=S01
RESOLUTION=2
WORKDIR=/public_bme2/bme-zhanghan/Jiayifan/dMRI/work
# QSIPrep call
singularity run -e \
-B $bids_dir/derivatives:/derivatives:ro \
-B $bids_dir/derivatives:/out \
-B ${WORKDIR}:${WORKDIR} \
-B /public_bme2/bme-zhanghan/Jiayifan/license.txt:/opt/freesurfer/license.txt \
/public_bme2/bme-zhanghan/Jiayifan/dMRI/qsiprep-1.0.0.sif \
/derivatives/nii /out participant \
--fs-license-file /opt/freesurfer/license.txt \
--output-resolution ${RESOLUTION} \
--work-dir ${WORKDIR} \
-v -v
But it didn’t run correctly and reported the following error:
INFO: Mounting image with FUSE.
e[91mERROR : Failed to create user namespace: user namespace disabled
e[0m
I also tried to refer to the posts of other experienced QSIPrep users I could find (Using QSIprep on Cheaha - Civitan International Neuroimaging Laboratory) and the modified script is as follows:
···
cd /public_bme2/bme-zhanghan/Jiayifan/dMRI
# run qsiprep
singularity run qsiprep-1.0.0.sif --participant_label $participant \
--fs_license_file /public_bme2/bme-zhanghan/Jiayifan/license.txt \
-v \
--separate_all_dwis \
--output_resolution 1.2 \
--skip_bids_validation \
$bids_dir \
$bids_dir/derivatives \
participant
The error is the same as the result of the last script run.
I’m not sure what went wrong, very much looking forward to your reply!
Best,
YJia