Hello,
First of all, thank you in advance for your time and feedback.
I am working with rhesus monkey brain data using NMT macaque template(9.1.2. Macaque template: NMT v2 (current) — AFNI, SUMA and FATCAT: v22.1.00).
I successfully created brain surfaces of various monkey subjects using the template.
Brain surfaces: inflated, sphere, white, pial, and etc.
With created surfaces, I am trying to register template CHARM(cortical regions) and SAMR(subcortical regions) mask volume (nii.gz file) using surface registration.
- I first created a surface overlay file using mri_vol2surf command.
mri_vol2surf --src ./CHARM_1_in_NMT_v2.0_sym.nii.gz --out lh.charm1.mgh --srcreg template.lta --hemi lh
-
Then, I used mri_surf2surf using template brain surface(NMT) and subject brain surface(let’s say Aida’s brain surfaces) to do surface to surface registration of the overlay file.
-
After that, I tried to make the warped(template->subject(Aida)) surface overlay file to a volume(nii.gz) with the function mri_surf2vol.
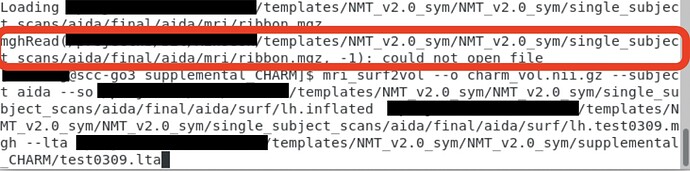
However, my current pipeline does not provide the ribbon.mgz file and the function fails.
→ Red box is the error
→ Commands under the red box is the command I used (mri_surf2vol) -
I am unable to create the ribbon.mgh file, because my pipeline does not provide other prerequisite files for creating ribbon.mgh file.
Would there be a way to resolve this issue
Or would there be another way to register template ROI volume mask to subject space using surface registration?
I am struggling with this issue for weeks.
Any help or feedback will be greatly appreciated.
Thank you so much.