TLDR:
Replicate the behavior of the SPM12 (v7771) “Segmentation” step with nipype in order to output a deformation field file y_*.nii
I would like to create a nipype tutorial that follows the preprocessing workflow from the SPM12 manual (Note: There is already a script in the the documentation that does this, but this follows the SPM8 manual, I am however using SPM12: v7771). I was already able to implement the first steps (Realignment & Coregistration). But I struggle with implementing the Segmentation step.
1.) First of all, there are multiple segmentation functions implemented and I don’t know exactly which of those replicate the behavior of SPM12 (v7771):
nipype.interfaces.spm.Segmentnipype.interfaces.spm.NewSegmentnipype.interfaces.spm.Segment
I think it is nipype.interfaces.spm.NewSegment?:
2.) I would like to follow the Segmentation step from the SPM12 manual (p. 235) which includes 3 steps:
Press the Segment button. This will call up the specification of a segmentation job in the batch
editor. Highlight the “Volumes” field and then select the subject’s registered anatomical image eg.
sM00223_002.img. Highlight “Save Bias Corrected” and select “Save Bias Corrected”. Highlight
“Deformation Fields” the bottom of the list and select “Forward”. Save the job file as segment.mat
and then press RUN. SPM will segment the structural image using the default tissue probability
maps as priors [9].
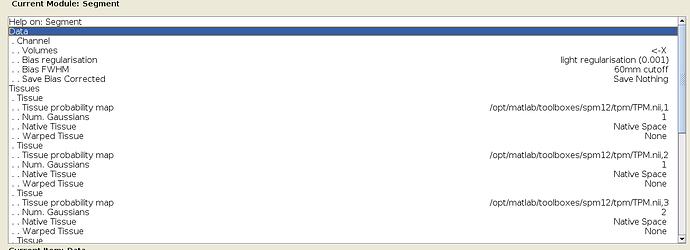
What are the minimum steps to reproduce these three steps when using nipype? When opening SPM12 on my machine, I see that a lot of stuff has default values, that I can just leave unchanged:
But nipype does not seem stick to those default values? E.g. in @PeerHerholz tutorial, he sets all values for the “Tissues” field manually to replicate the SPM12 default values. Also in the docs the channel_info field is set to again replicate values that are already set as default in SPM12:
seg.inputs.channel_info = (0.0001, 60, (True, True))