Hello,
I have done postprocessing on my fmriprepped data using nilearn.image.clean_image. The command I’m using is: nilearn.image.clean_img(data,session=None,detrend=True,standardize=True,confounds=confounds_array,low_pass=0.08,high_pass=0.008,t_r=1.5,ensure_finite=True,mask_img=mask). I removed the first 6 volumes of the raw data, prior to fmriprep.
The first volume of my fully processed data looks different from the rest. For example, this is the first volume:
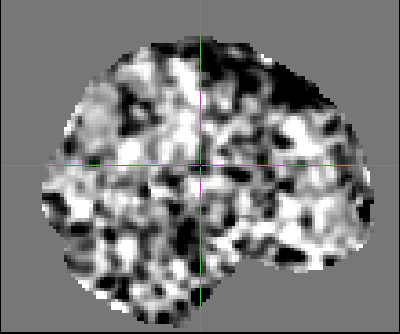
And this is the second volume:
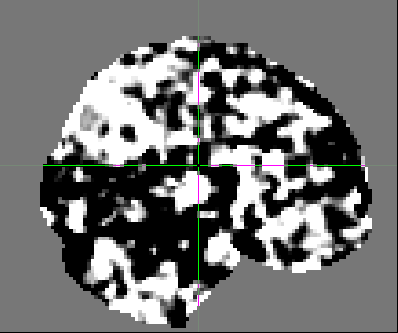
In addition, some of the participants data look quite different from the others. For example, this is another participant:
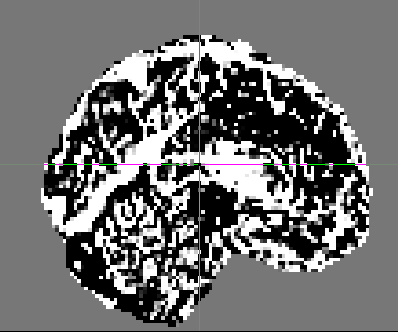
Do these data all look they way they should? Is it normal that they would look so different? Why is the first volume different than the rest for some people?
These images are not that weird. Maybe the question is whether the first image has dynamics (values range) different from the other. If you compute the variance in each image, is the first one strongly different from the others ?
It could simply be that the colormap you are using is not well-adapted.
My 2c,
Bertrand
Just to +1 this suggestion ! Also, depending on your scanner, you may need to discard several nonsteady-state or “dummy volumes” at the start of the scan, which will systematically have a different contrast than other acquired volumes.
fMRIPrep should detect these (and use them in BOLD reference generation, if available), but if you’re seeing consistent systematic differences between the variance of the first couple of volumes and subsequent volumes across participants, you might want to re-visit if fMRIPrep did in fact detect these correctly for your scans !
Hi. Did you ever figure out what the issue was? I am having a similar issue where the first and last volumes have a different value range when compared to the rest (even after discarding non steady state volumes).


