Dear NeroStars,
i am trying to re-sample a volume data (*nii.gz) to the surface, but not sure what is the most proper way to do it.
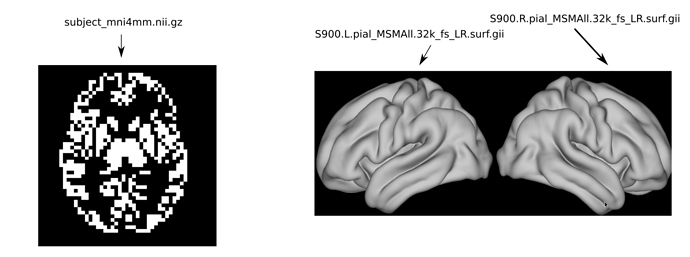
The volumetric data is a nifti file in MNI152 space, resolution of 4mm. The gray matter voxels are labeled there from 1 to 10k.
The target surface is the HCP_S900 surface, which has 32k vertices on each hemisphere (S900.L.pial_MSMAll.32k_fs_LR.surf.gii for the left hemisphere, and S900.R.pial_MSMAll.32k_fs_LR.surf.gii for the right hemisphere).
Now, I would like to register 10k gray matter voxels to the 64k surface. So far, I tried $wb_command -volume-to-surface-mapping, however, the surface-registered image did not really seem so nice. Is there any way to do the surface registration with freesurfer?
- the figure illustrates screenshot for the volume data and hcp_s900 surface templates.
Thanks in advance,
Seyma