Hi everyone,
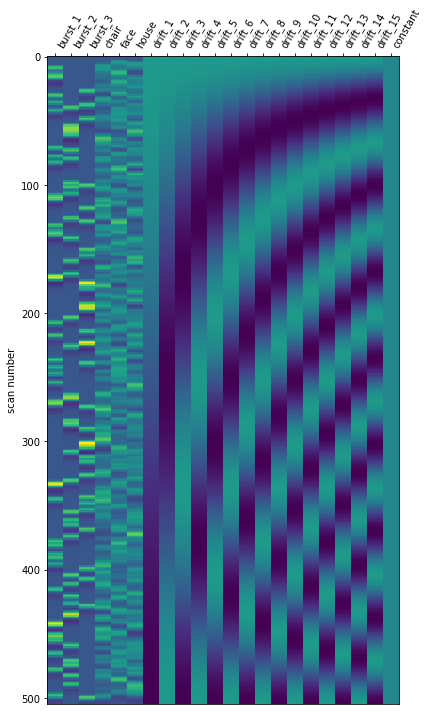
I am trying to run a 1st level analysis in nilearn. The experiment has 3 visual stimuli and 2 conditions. I have one design matrix per run, and each run has only one condition (like condition_1 for 1st and 2nd runs and condition_2 for 3rd and 4th runs). I need to:
- specify an F-contrast that will look at differences between stimulus_1 vs. stimulus_2 vs. stimulus_3 in one single contrast (rather than in 3 different pairwise contrasts);
- and specify a contrast comparing the 2 conditions (that, as said before, are in different runs).
However, I am having problems to specify those contrasts (and I’m new to fmri analysis so sorry if those questions are too basic). I checked some examples (like Simple example of two-runs fMRI model fitting and Single-subject data (two runs) in native space) but I am always getting errors when trying compute the contrasts (glm.first_level.FirstLevelModel.compute_contrast). Using 2D arrays always throw LinAlgError: Singular matrix and 1D contrasts crash when trying to broadcast the arrays (like ValueError: operands could not be broadcast together with remapped shapes [original->remapped]: (2,2) and requested shape (1,2)). Any clues on how to set those contrasts?
Thank you very much in advance!