Mentor: Daniele Marinazzo <daniele.marinazzo@gmail.com>
Skill level: Intermediate to Advanced
Required skills: Python, Docker; familiarity with neuroimaging tools (AFNI, FSL, FreeSurfer) would be beneficial
Time commitment: Full time (350 hours)
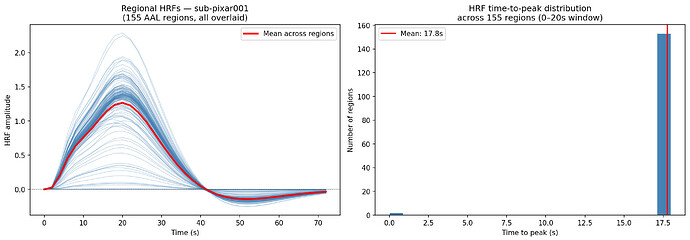
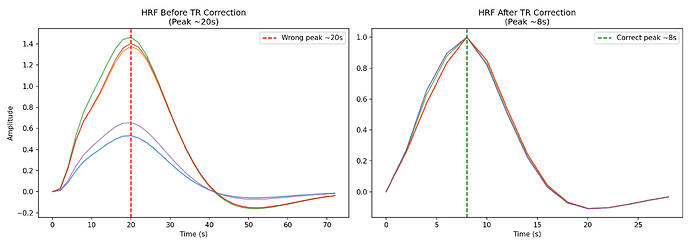
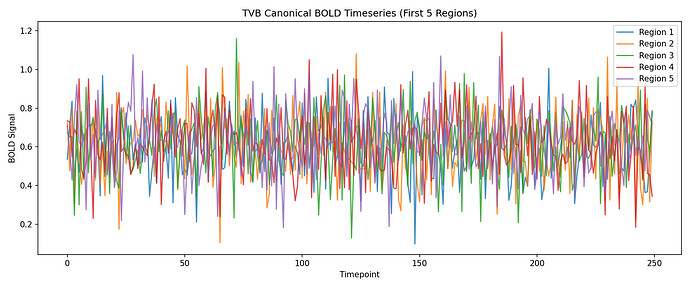
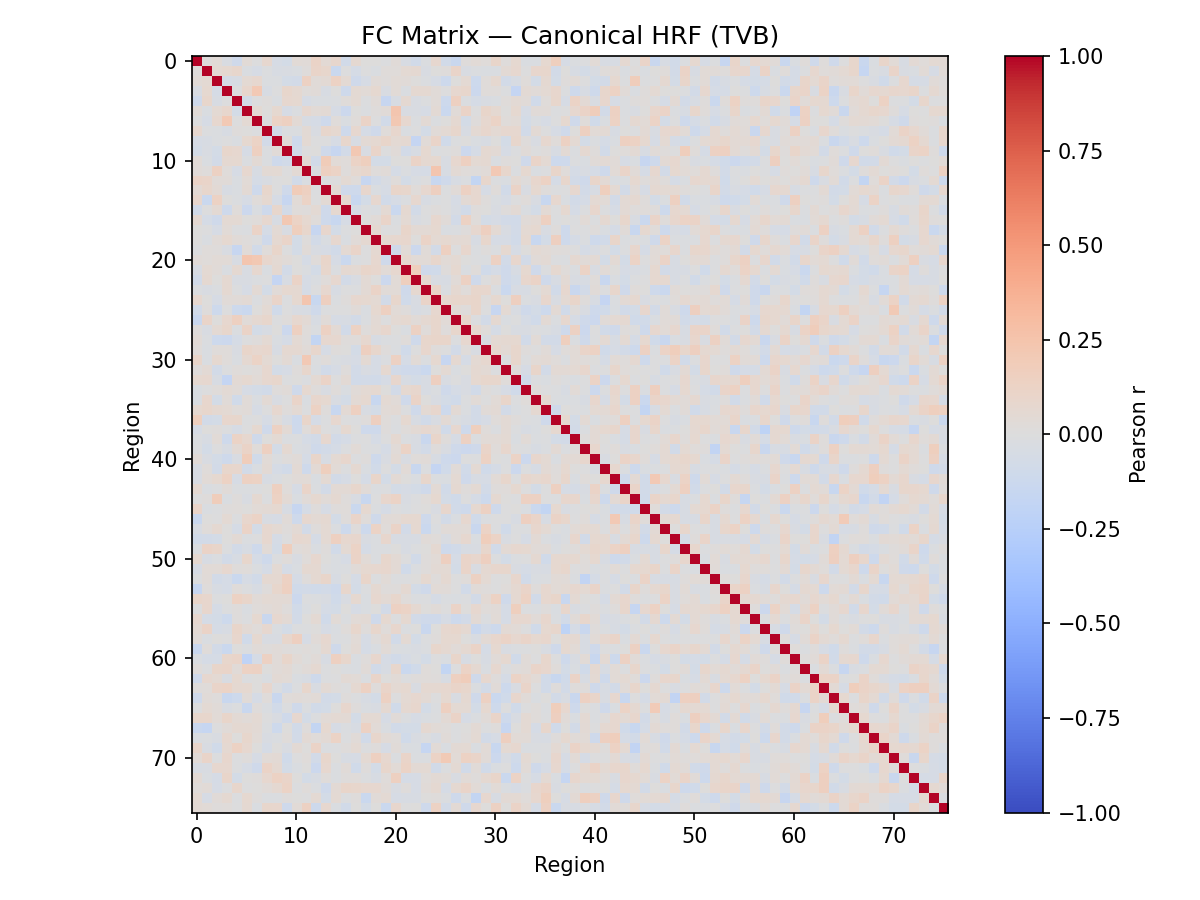
About: The hemodynamic response function (HRF) is a crucial mapping between neural activity and the recorded BOLD signal in fMRI experiments. Accounting for differences in the HRF shape is crucial when making inference on neural activity. The same should apply to computational models of large scale brain activity when the target is the BOLD signal or properties derived from it such as the Functional Connectivity. On the other hand models so far use a standard shape for the HRF. In resting state fMRI there is no explicit onset of neural activation, making it challenging to estimate the HRF using a linear model. A toolbox by our group proposes a point process model to estimate the HRF from resting state BOLD signals.
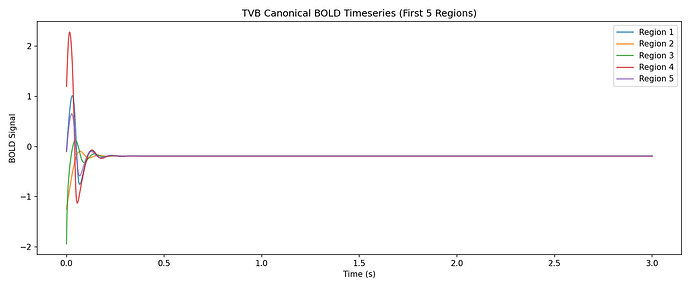
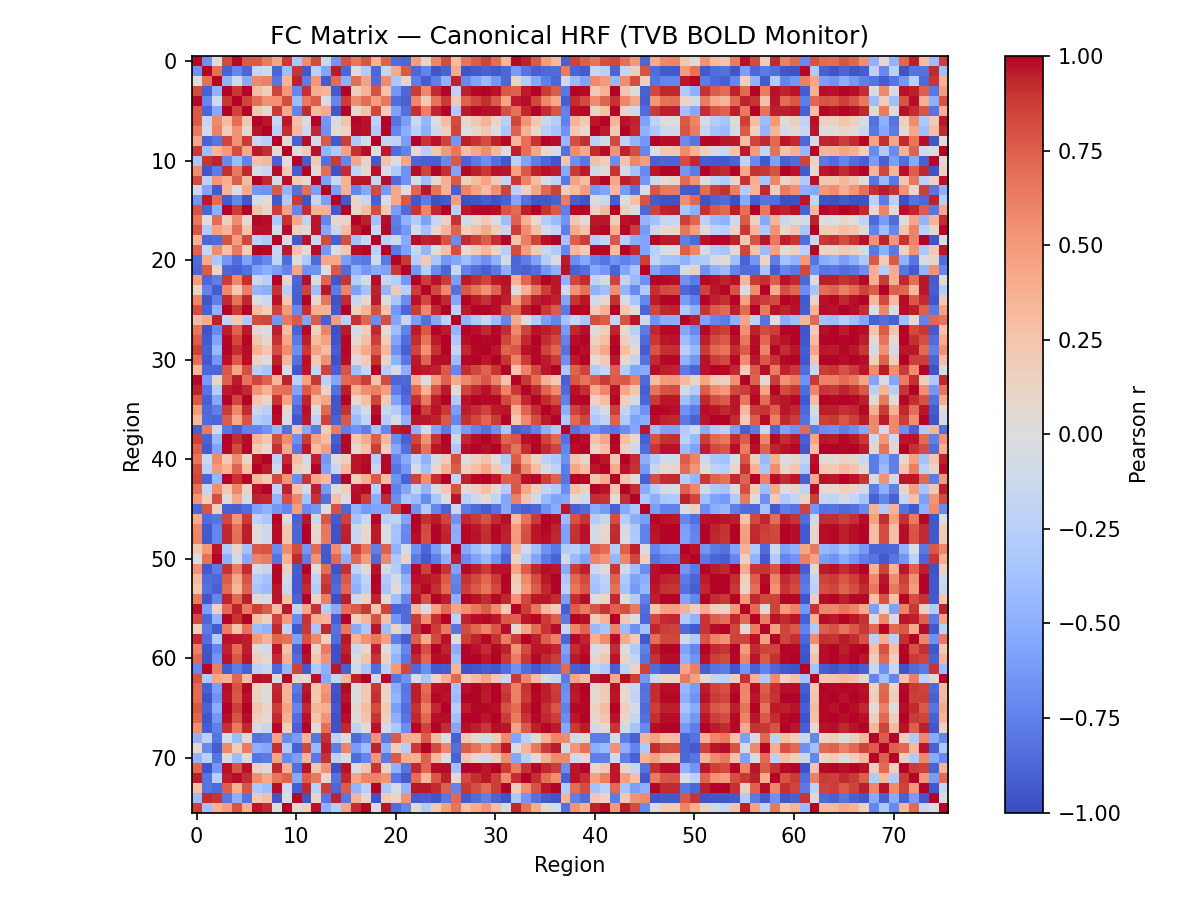
Aims: This project will have the aim to derive the empirical HRF for each brain region and each subject from BOLD recordings, and then to use this HRF in a computational model of brain activity (The Virtual Brain, implemented in EBRAINS). The legacy approach will be compared with the proposed one.
Website: rsHRF | EBRAINS
Tech keywords: Python, Docker, The Virtual Brain, rsHRF