Hi everyone,
I’m running heudiconv on singularity to convert DICOMs to BIDS. All MPRAGE and rest scans seems to have converted correctly, but I’m having error with only the topup scans.
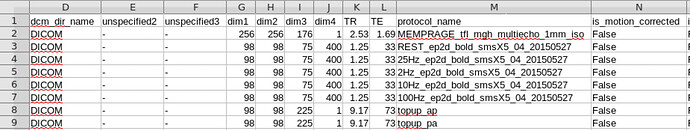
Here in dicominfo.tsv file, dimension 3 should be 75 and dimension 4 should be 3 in topup scans, but seems to be concatenating volumes.
Here’s the heuristic file:
import os
def create_key(template, outtype=(‘nii.gz’,), annotation_classes=None):
if template is None or not template:
raise ValueError(‘Template must be a valid format string’)
return template, outtype, annotation_classesdef infotodict(seqinfo):
“”“Heuristic evaluator for determining which runs belong where
allowed template fields - follow python string module:
item: index within category
subject: participant id
seqitem: run number during scanning
subindex: sub index within group
“””
t1w = create_key(‘sub-{subject}/{session}/anat/sub-{subject}_{session}_run-00{item:01d}T1w’)
func_rest = create_key('sub-{subject}/{session}/func/sub-{subject}{session}_task-rest_run-00{item:01d}bold’)
func_2Hz = create_key('sub-{subject}/{session}/func/sub-{subject}{session}_task-stim2Hz_run-00{item:01d}bold’)
func_10Hz = create_key('sub-{subject}/{session}/func/sub-{subject}{session}_task-stim10Hz_run-00{item:01d}bold’)
func_25Hz = create_key('sub-{subject}/{session}/func/sub-{subject}{session}_task-stim25Hz_run-00{item:01d}bold’)
func_100Hz = create_key('sub-{subject}/{session}/func/sub-{subject}{session}_task-stim100Hz_run-00{item:01d}bold’)
topup_ap = create_key('sub-{subject}/{session}/fmap/sub-{subject}{session}dir-AP_epi’)
topup_pa = create_key('sub-{subject}/{session}/fmap/sub-{subject}{session}_dir-PA_epi’)info = {t1w: [], func_rest: [], func_2Hz: [], func_10Hz: [], func_25Hz: [], func_100Hz: [], topup_ap: [], topup_pa: []} for idx, s in enumerate(seqinfo): if (s.dim1 == 256) and (s.dim2 == 256) and ('MEMPRAGE_tfl_mgh_multiecho_1mm_iso' in s.protocol_name): info[t1w].append(s.series_id) if (s.dim1 == 98) and (s.dim4 == 400) and ('REST_ep2d_bold_smsX5_04_20150527' in s.protocol_name): info[func_rest].append(s.series_id) if (s.dim1 == 98) and (s.dim4 == 400) and ('2Hz_ep2d_bold_smsX5_04_20150527' in s.protocol_name): info[func_2Hz].append(s.series_id) if (s.dim1 == 98) and (s.dim4 == 400) and ('10Hz_ep2d_bold_smsX5_04_20150527' in s.protocol_name): info[func_10Hz].append(s.series_id) if (s.dim1 == 98) and (s.dim4 == 400) and ('25Hz_ep2d_bold_smsX5_04_20150527' in s.protocol_name): info[func_25Hz].append(s.series_id) if (s.dim1 == 98) and (s.dim4 == 400) and ('100Hz_ep2d_bold_smsX5_04_20150527' in s.protocol_name): info[func_100Hz].append(s.series_id) if (s.dim1 == 98) and (s.dim2 == 98) and ('topup_ap' in s.protocol_name): info[topup_ap].append(s.series_id) if (s.dim1 == 98) and (s.dim2 == 98) and ('topup_pa' in s.protocol_name): info[topup_pa].append(s.series_id) return info
Could this error be based on the initial DICOM files?
Do you have any idea what is happening?
Thanks a lot!
Kyungsun