Hi friends,
following issue: I’m following this tutorial to convert .dcm to BIDS: BIDS Tutorial Series: HeuDiConv Walkthrough | Stanford Center for Reproducible Neuroscience. I’m stuck on step 4, where I run HeuDiConv with heuristic.py:
docker run --rm -it -v C:/Users/pazlab/Documents/Max/MRIData/Pilot:/base nipy/heudiconv:latest -d /base/{subject}/*/*/*/*.dcm -o /base/Nifti/ -f /base/Nifti/code/heuristic.py -s "BE796" -c dcm2niix -b --overwrite
Running this, I get the following output:
C:\Users\pazlab>docker run --rm -it -v C:/Users/pazlab/Documents/Max/MRIData/Pilot:/base nipy/heudiconv:latest -d /base/{subject}////.dcm -o /base/Nifti/ -f /base/Nifti/code/heuristic.py -s “BE796” -c dcm2niix -b --overwrite
INFO: Running heudiconv version 0.9.0 latest 0.10.0
INFO: Need to process 1 study sessions
INFO: PROCESSING STARTS: {‘subject’: ‘BE796’, ‘outdir’: ‘/base/Nifti/’, ‘session’: None}
INFO: Processing 2104 dicoms
INFO: Analyzing 2104 dicoms
INFO: Generated sequence info for 14 studies with 2104 entries total
INFO: Doing conversion using dcm2niix
INFO: Lock 140494038703856 acquired on /base/Nifti/heudiconv.lock
INFO: Populating template files under /base/Nifti/
INFO: Lock 140494038703856 released on /base/Nifti/heudiconv.lock
INFO: PROCESSING DONE: {‘subject’: ‘BE796’, ‘outdir’: ‘/base/Nifti/’, ‘session’: None}
And in my Nifti folder, I get no .json outputs except my participants .json.
It looks like it’s not recognizing the identifiers I give in my dicominfo.tsv file but I’m not sure why, or at least for some other reason it seems like it doesn"t manage to read the files.
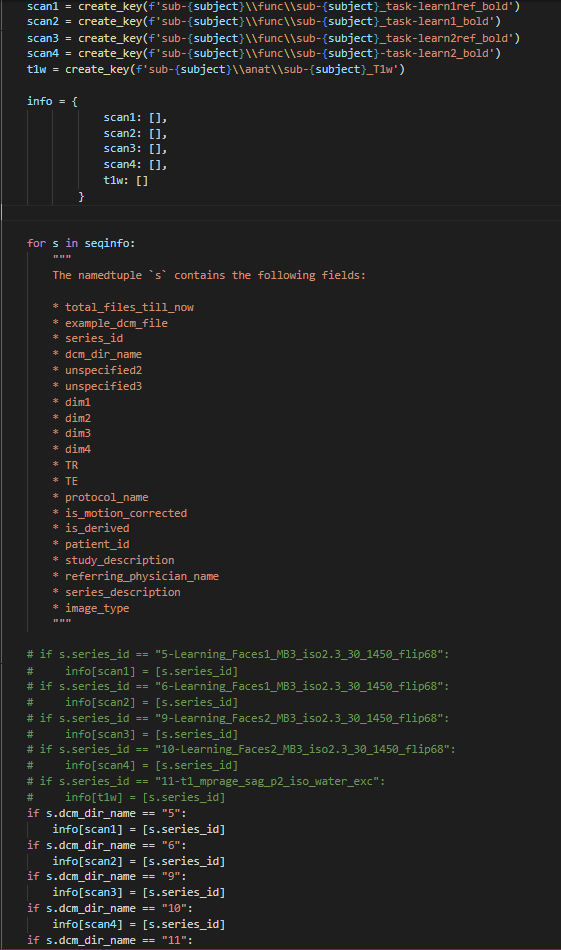
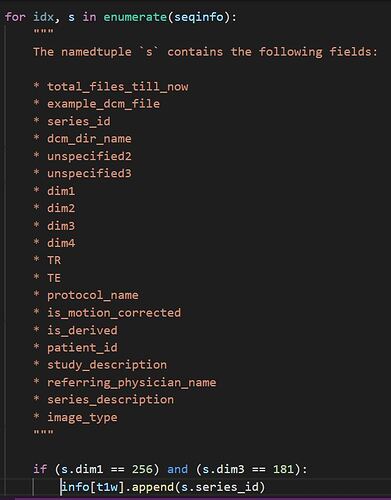
Here’s my code:
I’ve tried lots of different things, among them:
- Changed the file identifier (as you can see in the bracketed out part)
- Playing around with syntax (i.e. different backslashes etc., as you can see on top of the picture)
Not sure exactly what the issue is. Any leads welcome!
Thanks
Max