Hello World,
I am trying to perform a beta series analysis on fMRIPREP’d data. Therefore, I need my mask to be in the same space as my functional data. My functional data is in MNI_asym space: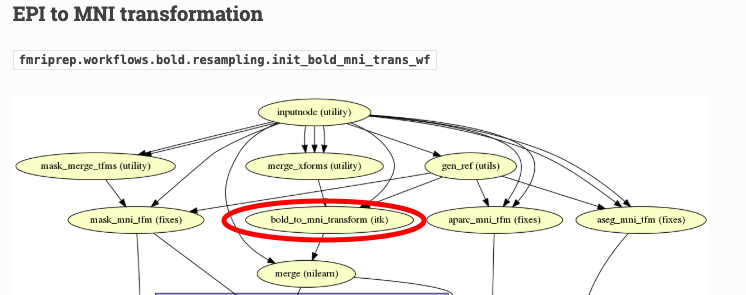
So I need the affine transform matrix so I can align the mask I have to the MNI space. In FEAT this file would be reg/example_func2standard.mat. However, I cannot find the transformation in fMRIPREP.
Overall this is what I am trying to accomplish:
flirt -in my_mask.nii.gz -ref EPIscan.nii.gz -init EPI2standard.mat -applyxfm -out my_mask_standard.nii.gz
fslmaths my_mask_standard -thr 0.9 -bin my_mask_standard_bin
based off https://www.jiscmail.ac.uk/cgi-bin/webadmin?A2=FSL;f63f806d.1007
Thanks!
Just to clarify, your mask is currently in the subject’s native BOLD/EPI space, and you want to align it to MNI asym space?
To transform data from native space to MNI space, fMRIPrep uses a vector field deformation computed by composing (1) the affine transform between the subject’s BOLD space and the subject’s anatomical space and (2) the vector-field transform from the subject’s anatomical space to the standard space (in this case, MNI). Thus, you won’t find an BOLD-to-MNI affine transform matrix among the outputs: this would yield a poor alignment in comparison with fMRIPrep's nonlinear approach.
If your mask is currently in the subject’s native BOLD space, I would instead recommend moving it to MNI space by composing transforms (1) and (2) above.
(1) is <identifiers>_from-orig_to-T1w_mode-image_xfm.txt
(2) is <identifiers>_from-T1w_to-MNI152NLin2009cAsym_mode-image_xfm.h5
You can compose and apply them using antsApplyTransforms to warp the mask into MNI space as follows:
antsApplyTransforms \
-i </path/to/mask.nii.gz> \
-o </path/to/mask_space-MNI152NLin2009cAsym.nii.gz> \
-r </path/to/reference_MNI_template.nii.gz> \
-n NearestNeighbor \
-t <identifiers>_from-T1w_to-MNI152NLin2009cAsym_mode-image_xfm.h5 \
-t <identifiers>_from-orig_to-T1w_mode-image_xfm.txt
Note that the above call would change if your mask isn’t binary-valued.
1 Like
@rastko I am wondering if I can get EPI2standard.mat for another purpose, that is, to perform ICA-AROMA.
Is it possible to create the mat file from xxx_from-orig_to-T1w_mode-image_xfm.txt?
I would deeply appreciate it if you could show how to do this.
Thanks in advance!
