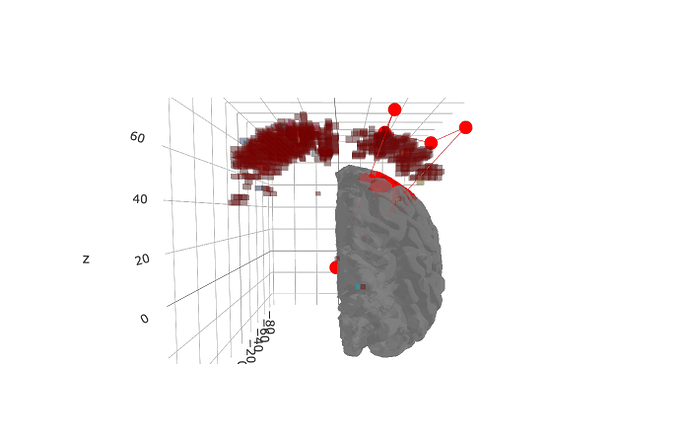
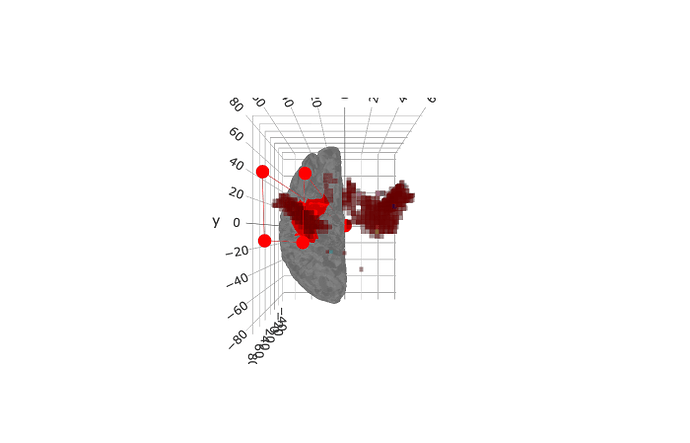
I am trying to resample volume to lh.pial to align fmap and mri surface and have an issues, shown on screenshot.
map is higher than freesurfer surface.
mri_info of both volume and pial are:
fmap volume:
ras xform present
xform info: x_r = 1.0000, y_r = 0.0000, z_r = -0.0000, c_r = 3.3750
: x_a = 0.0000, y_a = 1.0000, z_a = -0.0000, c_a = -15.7500
: x_s = 0.0000, y_s = 0.0000, z_s = 1.0000, c_s = 19.5000
Orientation : RAS
pial surface:
('xras', array([-1., 0., 0.])),
('yras', array([ 0., 0., -1.])),
('zras', array([0., 1., 0.])),
('cras', array([-3.433, 18.419, 28.598]))]))
how resample one of them to the center of coordinates? I suppose i might use xform somehow? Found similar problem in https://github.com/nipy/nibabel/issues/518, @oesteban, Have you solved it?