Thank you! I think your diagnosis of the problem as due to the poor fmap is spot on. This participant was in in two waves of scanning, three sessions each wave. I pulled the other five pairs of fmaps and added them to the shared box folder.
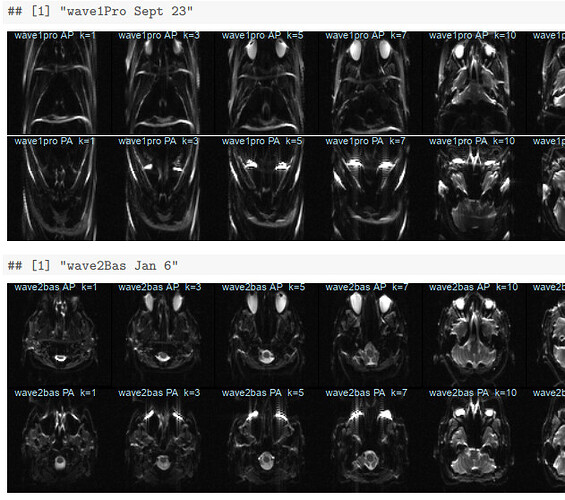
I plotted slices from each of them in order of session date in a knitr pdf now in the root of that box directory. This person’s sessions were closer together in time than many of our participants; all six sessions were collected over less than six months. The two sessions from late September (wave1rea and wave1pro) have clearly more distortion in the fmaps than the rest:
Looking at these, it’s surprising we didn’t have more issues with preprocessing these sessions! My QC procedure has been just checking that the fmaps have brains; clearly I should look more closely. The distortion in the two late September sessions seems similar; suggesting this was a scanner problem, rather than something like movement during the fmap acquisitions? I can check if we collected any other participants during this time frame, and if so, how their fmaps look.
Some of the wave1rea runs are currently running through fmriprep 21.0.1; I’ll share how it turns out when we have results.