Hi @mblesac ,
Which fmriprep version are you working with? As you must have noted (and perhaps wrote about it already) there are several posts covering this issue with SDC not working well in FMRIPREP, especially for the “Direct Fieldmap” approach (Two magnitude and one fiedlmap images). Some corrections were made for version 23.0.0, perhaps you could try it?
As it was rightfully stated by @schillkg , synbold-disco was designed for cases where no fieldmap were acquired: normally if all goes well, SDC should work at least as well as synbold-disco for bold images for which proper fieldmap were acquired.
Anyway, if you want to give a try to the synbold-disco/fmriprep method for you dataset, for the method that worked correctly in my test, here are the steps I followed:
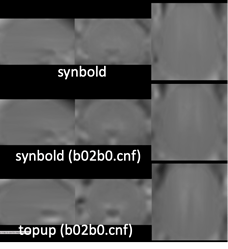
1/ run synbold-disco for your distorted bold serie. You must rename your bold image as: BOLD_d.nii.gz and your T1w image as: T1.nii.gz and place those in the INPUTS directory. Then run synbold-disco. In my case I used singularity with the following command:
singularity run -e \
-B /scratch/jsein/BIDS/$study/sourcedata/synbold/sub-${sub}/INPUTS:/INPUTS \
-B /scratch/jsein/BIDS/$study/derivatives/synbold/sub-${sub}:/OUTPUTS \
-B /scratch/jsein/BIDS/freesurfer/license.txt:/opt/freesurfer/license.txt \
/scratch/jsein/my_images/synbold-disco-v1.3.simg --stripped --motion_corrected
2/ For the moment, synbold-disco doesn’t call topup with the --fout option to output the fieldmap in Hz. So I proceeded by calling topup once more on the OUTPUTS of synbold-disco, through singularity again, using my edited acqparamsJS.txt file:
cat acqparamsJS.txt
0 1 0 real_total_readout_time_of_bold # my bold image are with PE direction in P->A
0 1 0 0
singularity exec -e -B /scratch/jsein/BIDS/$study \
/scratch/jsein/my_images/fmriprep-23.0.0.simg \
topup --imain=/scratch/jsein/BIDS/$study/derivatives/synbold/sub-${sub}/BOLD_all.nii.gz \
--datain=/scratch/jsein/BIDS/$study/derivatives/synbold/sub-${sub}/acqparamsJS.txt \
--config=b02b0.cnf --subsamp=1,1,1,1,1,1,1,1,1 --miter=10,10,10,10,10,20,20,30,30 \
--lambda=0.00033,0.000067,0.0000067,0.000001,0.00000033,0.000000033,0.0000000033,0.000000000033,0.00000000000067 --scale=0 \
--out=/scratch/jsein/BIDS/$study/derivatives/synbold/sub-${sub}/my_topup_results \
--fout=/scratch/jsein/BIDS/$study/derivatives/synbold/sub-${sub}/my_field \
--iout=/scratch/jsein/BIDS/$study/derivatives/synbold/sub-${sub}/my_unwarped_images \
--jacout=jac --rbmout=xfm --dfout=warpfield -v
3/ Rename the files needed for SDC within FMRIPREP according the “direct fieldmap” method:
-
my_field.nii.gz(created in step 2/) → fmap/sub-SUB_ses-SES_fieldmap.nii.gz
- copy the JSON file from your func image into
fmap/sub-SUB_ses-SES_fieldmap.jsonand add the following tags:
"Units": "Hz",
"IntendedFor": [
"ses-SES/func/sub-SUB_ses-SES_task-TASK_bold.nii.gz"
]
(you could also use the B0FieldIdentifier / B0fieldSource strategy).
-
BOLD_s_3D.nii.gz → fmap/sub-SUB_ses-SES_magnitude.nii.gz
Now you are all set to run FMRIPREP using the fieldmap estimated the synbold-disco!
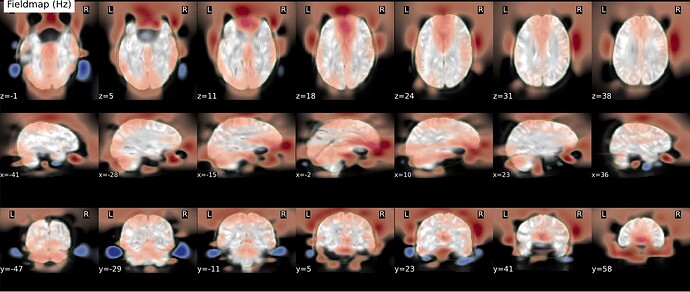
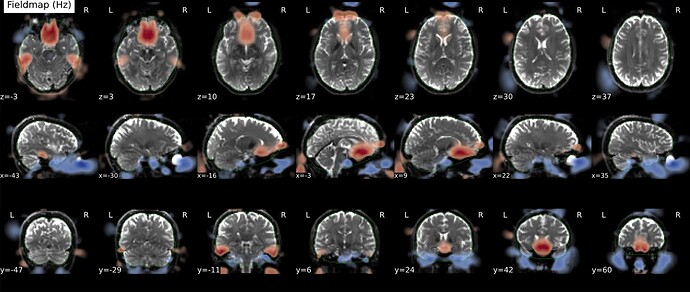
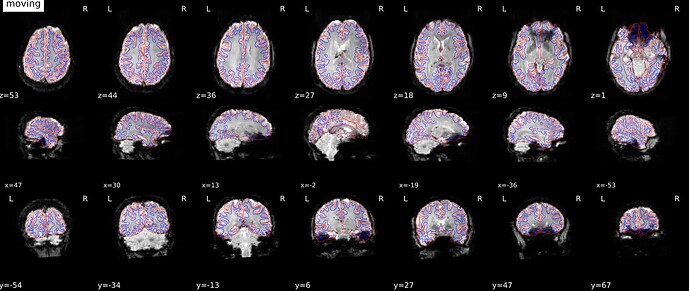
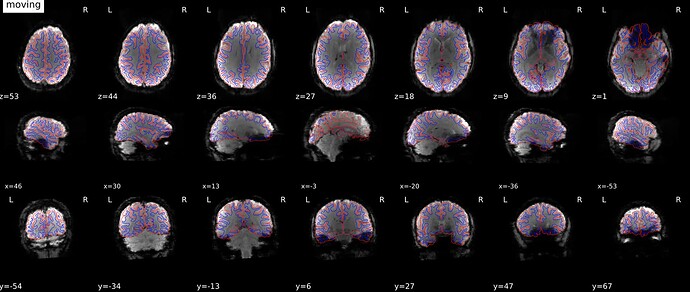
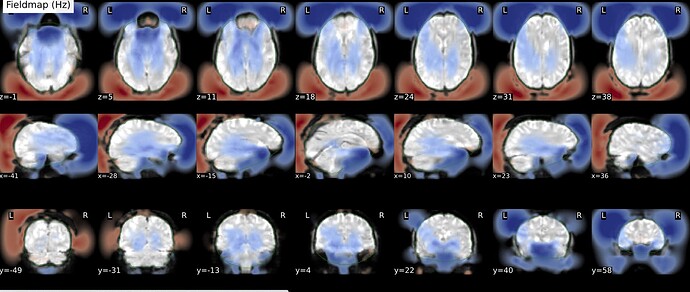
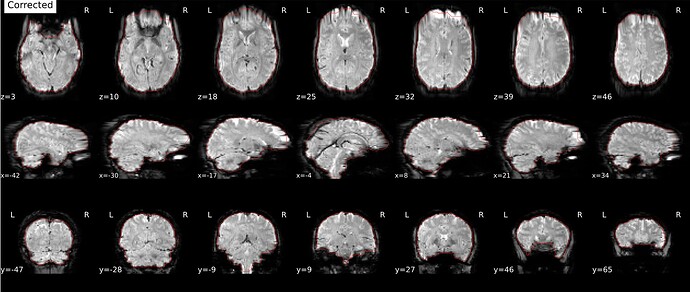
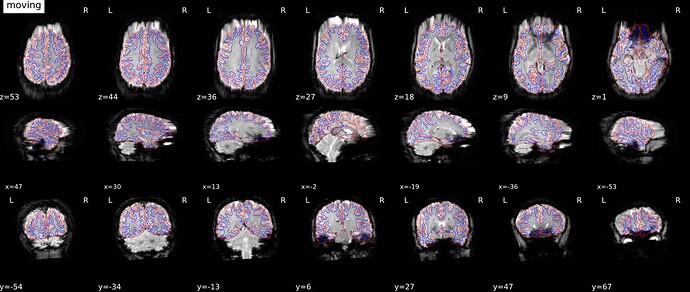
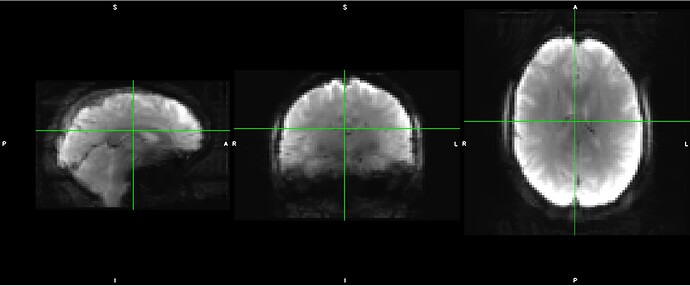
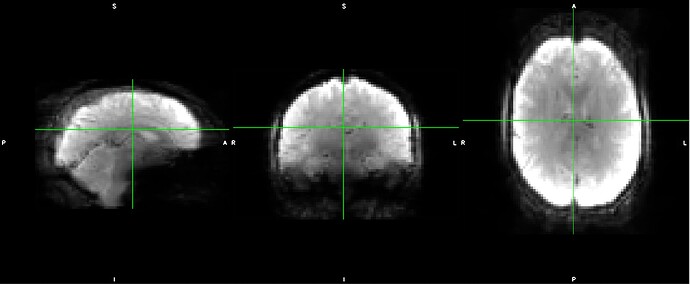
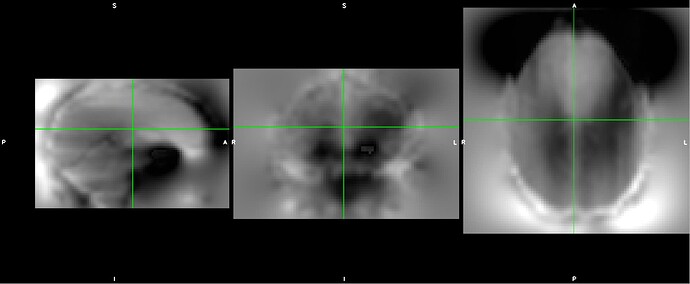
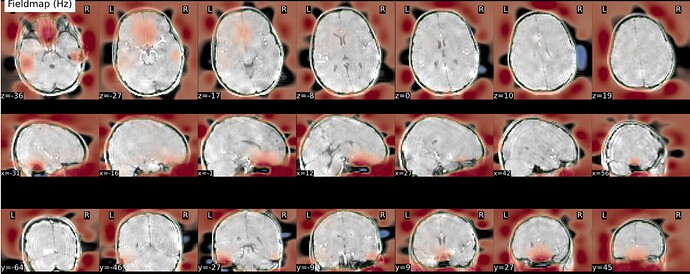
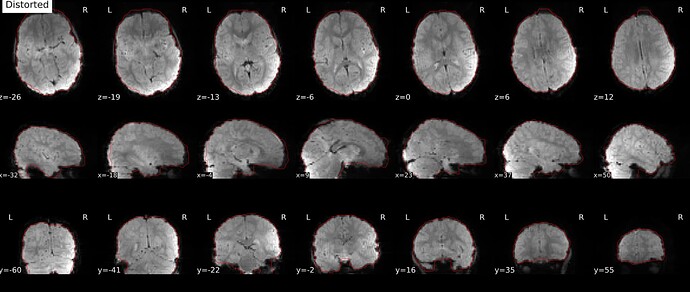
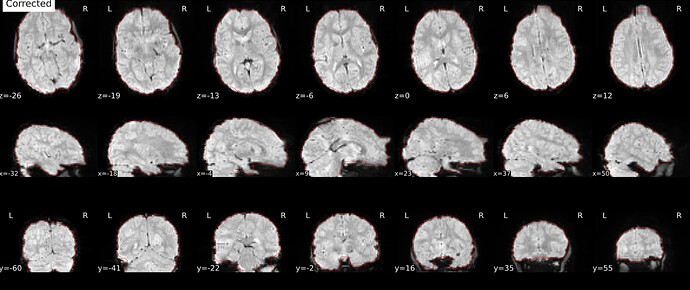
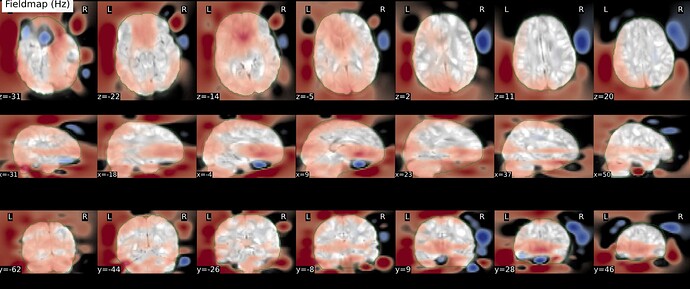
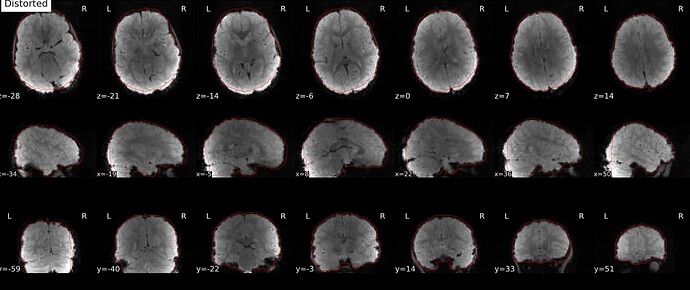
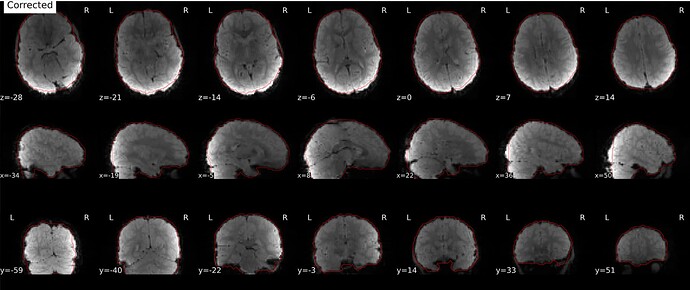
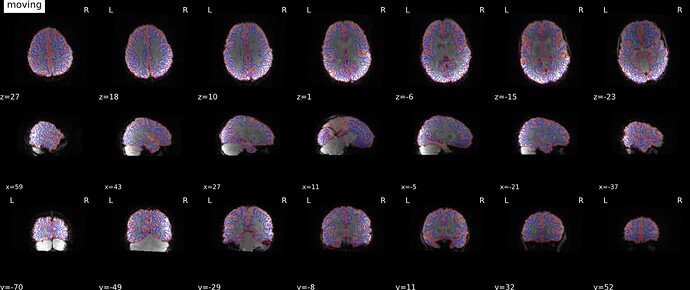
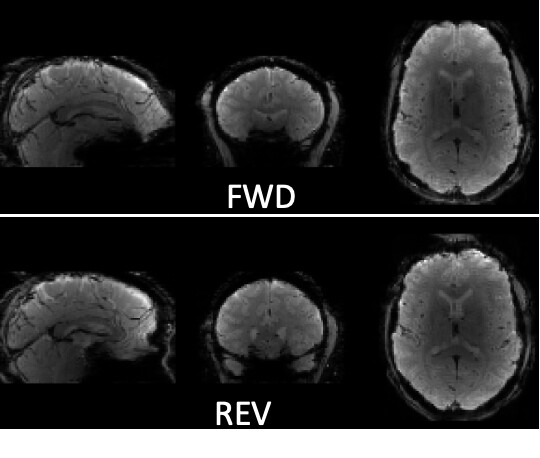
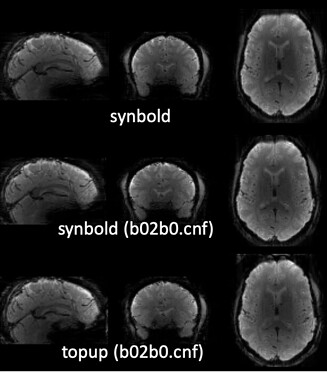
As stated above, the fieldmap calculated by synbold-disco has a strange appearance but the correction looked correct (at least for the one subject I tried so far).
I can’t guaranty it will work for you but it would be interesting to see if you get good results for your dataset with this method.