Hi @effigies, sorry for me being impatient!
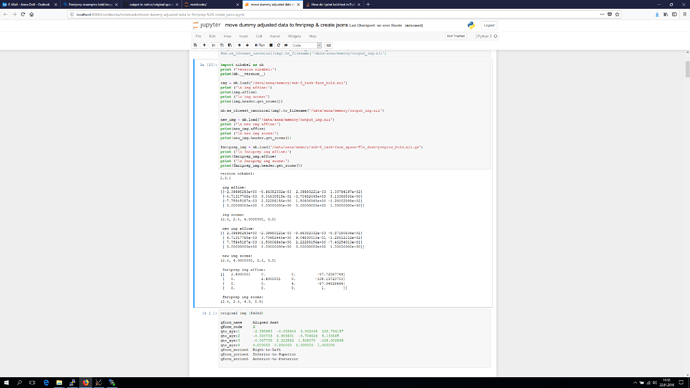
I doublechecked but still sent you the information of the T1 instead of the original bold image, sorry. Below you see a screenshot of the python code you suggested, and as a last entry I added the fslhd info, this time really of the original bold image.
I’m using 6 dof BOLD T1w registration according to the html report. Further we just noticed " Note on orientation: qform matrix overwritten. The qform has been copied from sform." -> is this of importance?
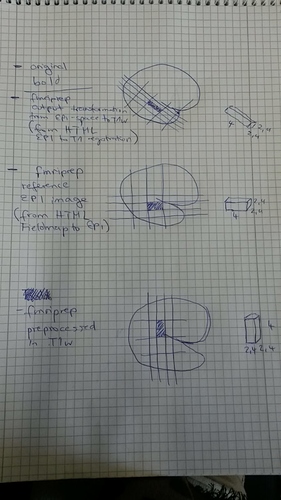
Below a try to summarize the different voxel orientation in the different images:
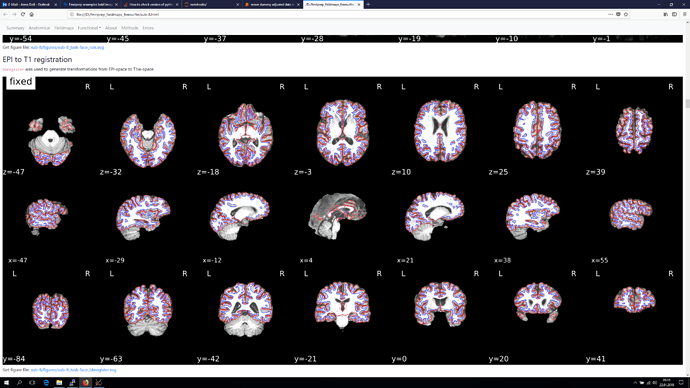
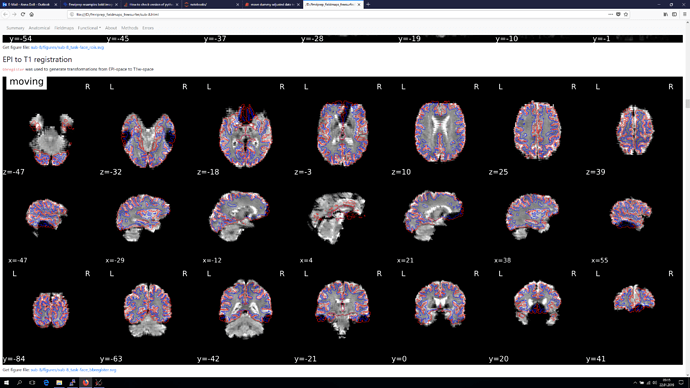
The registration section of the html reports
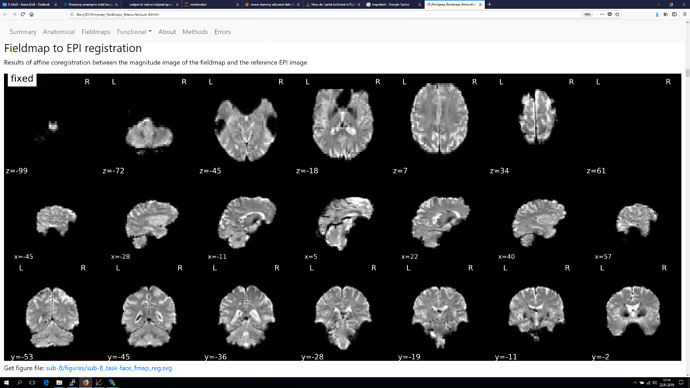
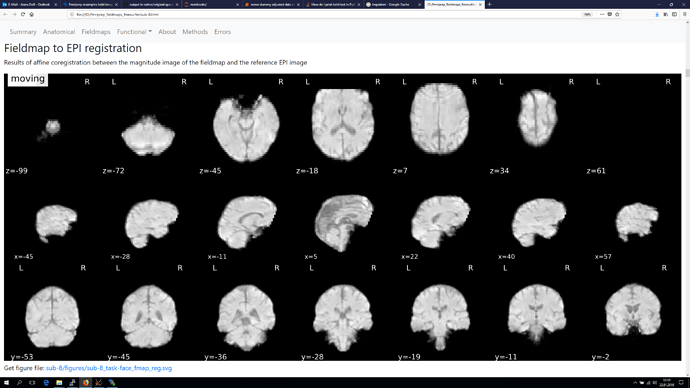
and finally also screenshots from the Fieldmap to EPI registration, as orientation of the voxel seems to be different again in those.
Sharing a T1w image and volumes of the BOLD series of a healthy control would be possible. I try to get the data today from the MRI just in case. Those bold images are oriented coronal along the hippocampus. The axial slices I refered to are from a different bold series of our team (you find them on open neuro https://openneuro.org/datasets/ds001419/versions/1.0.1 ).
I hope I did not miss anything and those information helps to understand what is happening. Thank you!!